α-tubulin detyrosination fine-tunes kinetochore-microtubule attachments
- PMID: 39521805
- PMCID: PMC11550433
- DOI: 10.1038/s41467-024-54155-8
α-tubulin detyrosination fine-tunes kinetochore-microtubule attachments
Abstract
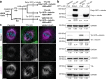
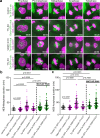
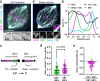
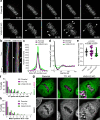
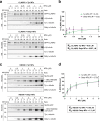
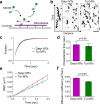
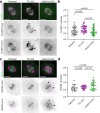
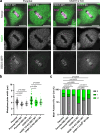
Post-translational cycles of α-tubulin detyrosination and tyrosination generate microtubule diversity, the cellular functions of which remain largely unknown. Here we show that α-tubulin detyrosination regulates kinetochore-microtubule attachments to ensure normal chromosome oscillations and timely anaphase onset during mitosis. Remarkably, detyrosinated α-tubulin levels near kinetochore microtubule plus-ends depend on the direction of chromosome motion during metaphase. Proteomic analyses unveil that the KNL-1/MIS12/NDC80 (KMN) network that forms the core microtubule-binding site at kinetochores and the microtubule-rescue protein CLASP2 are enriched on tyrosinated and detyrosinated microtubules during mitosis, respectively. α-tubulin detyrosination enhances CLASP2 binding and NDC80 complex diffusion along the microtubule lattice in vitro. Rescue experiments overexpressing NDC80, including variants with slower microtubule diffusion, suggest a functional interplay with α-tubulin detyrosination for the establishment of a labile kinetochore-microtubule interface. These results offer a mechanistic explanation for how different detyrosinated α-tubulin levels near kinetochore microtubule plus-ends fine-tune load-bearing attachments to both growing and shrinking microtubules.
© 2024. The Author(s).
Conflict of interest statement
The authors declare no competing interests.
Figures

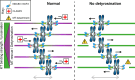
References
Publication types
MeSH terms
Substances
Grants and funding
- 681443/EC | EU Framework Programme for Research and Innovation H2020 | H2020 Priority Excellent Science | H2020 European Research Council (H2020 Excellent Science - European Research Council)
- 2021.07945.BD/Ministry of Education and Science | Fundação para a Ciência e a Tecnologia (Portuguese Science and Technology Foundation)
- PTDC/MED-ONC/3479/2020/Ministry of Education and Science | Fundação para a Ciência e a Tecnologia (Portuguese Science and Technology Foundation)
- LCF/PR/HR21/52410025/"la Caixa" Foundation (Caixa Foundation)
- R35 GM141747/GM/NIGMS NIH HHS/United States
- R35-GM141747/U.S. Department of Health & Human Services | National Institutes of Health (NIH)
- SFRH/BD/141066/2018/Ministry of Education and Science | Fundação para a Ciência e a Tecnologia (Portuguese Science and Technology Foundation)
- SFRH/BD/52044/2012/Ministry of Education and Science | Fundação para a Ciência e a Tecnologia (Portuguese Science and Technology Foundation)
LinkOut - more resources
Full Text Sources

