TNO155 is a selective SHP2 inhibitor to target PTPN11-dependent oral squamous cell carcinoma
- PMID: 39524880
- PMCID: PMC11550046
- DOI: 10.1016/j.heliyon.2024.e39677
TNO155 is a selective SHP2 inhibitor to target PTPN11-dependent oral squamous cell carcinoma
Abstract
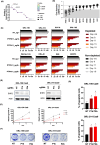
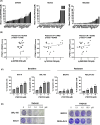
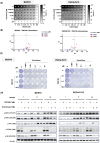
Oral squamous cell carcinoma (OSCC) is known to be driven by multiple intricated receptor tyrosine kinases (RTKs) including EGFR, PI3K/AKT and MAPK signaling pathways. However, whilst targeting EGFR with cetuximab has been approved for the treatment of OSCC, other single-agent inhibitors of the RTKs have shown modest effects in improving survival. From the genome-wide CRISPR/Cas9 screen on 21 OSCC cell lines, we have identified PTPN11 among the top essential genes in OSCC. PTPN11 encodes for SHP2, a phosphatase that acts as a master signal transducer, downstream of various RTKs. Although PTPN11 overexpression has been reported in OSCC, little is known about its role as an essential gene for OSCC survival and its potential as a therapeutic target. Herein, we confirmed that PTPN11 is an essential gene in OSCC where its deletion significantly impacted cell survival. We evaluated three SHP2 inhibitors on 21 OSCC cell lines and found TNO155 to be significantly associated with CRISPR dependency score. We showed that TNO155 caused dose-dependent suppression on p-ERK and p-MEK, and suppresses the JAK/STAT pathway via downregulating p-JAK1, p-STAT1, p-STAT3. Furthermore, we confirmed that the combination of the mTOR inhibitor, everolimus with TNO155 is synergistic in OSCC. In summary, PTPN11 is a promising therapeutic target in OSCC that can be selectively targeted by SHP2 inhibitor such as TNO155. Our findings on the use of mTOR inhibitor, everolimus to overcome resistance to TNO155 are essential to inform on next phases of clinical trials which is warranted for the treatment of OSCC.
Keywords: Allosteric inhibitor; Drug resistance; Essential genes; OSCC; PTPN11; Protein tyrosine phosphatase; RTK signaling; SHP2.
© 2024 The Authors.
Figures

Similar articles
-
Combinations with Allosteric SHP2 Inhibitor TNO155 to Block Receptor Tyrosine Kinase Signaling.Clin Cancer Res. 2021 Jan 1;27(1):342-354. doi: 10.1158/1078-0432.CCR-20-2718. Epub 2020 Oct 12. Clin Cancer Res. 2021. PMID: 33046519
-
Protein Tyrosine Phosphatase Non-Receptor 11 (PTPN11/Shp2) as a Driver Oncogene and a Novel Therapeutic Target in Non-Small Cell Lung Cancer (NSCLC).Int J Mol Sci. 2023 Jun 23;24(13):10545. doi: 10.3390/ijms241310545. Int J Mol Sci. 2023. PMID: 37445722 Free PMC article.
-
MicroRNA-186 serves as a tumor suppressor in oral squamous cell carcinoma by negatively regulating the protein tyrosine phosphatase SHP2 expression.Arch Oral Biol. 2018 May;89:20-25. doi: 10.1016/j.archoralbio.2018.01.016. Epub 2018 Jan 31. Arch Oral Biol. 2018. PMID: 29407635
-
Tyrosine phosphatase PTPN11/SHP2 in solid tumors - bull's eye for targeted therapy?Front Immunol. 2024 Mar 5;15:1340726. doi: 10.3389/fimmu.2024.1340726. eCollection 2024. Front Immunol. 2024. PMID: 38504984 Free PMC article. Review.
-
Strategies to overcome drug resistance using SHP2 inhibitors.Acta Pharm Sin B. 2021 Dec;11(12):3908-3924. doi: 10.1016/j.apsb.2021.03.037. Epub 2021 Mar 28. Acta Pharm Sin B. 2021. PMID: 35024315 Free PMC article. Review.
Cited by
-
Implication of protein post translational modifications in gastric cancer.Front Cell Dev Biol. 2025 Feb 4;13:1523958. doi: 10.3389/fcell.2025.1523958. eCollection 2025. Front Cell Dev Biol. 2025. PMID: 39968176 Free PMC article. Review.
-
Novel receptor tyrosine kinase-targeted strategies to overcome resistance in oral squamous cell carcinoma.Pharmacol Rep. 2025 Aug;77(4):962-982. doi: 10.1007/s43440-025-00745-2. Epub 2025 Jun 5. Pharmacol Rep. 2025. PMID: 40468099 Review.
References
-
- Torre L.A., et al. Global cancer statistics, 2012. CA A Cancer J. Clin. 2015;65(2):87–108. - PubMed
-
- J F., et al. International Agency for Research on Cancer; Lyon, France: 2020. Global Cancer Observatory: Cancer Today.
-
- Chai A.W.Y., Lim K.P., Cheong S.C. Translational genomics and recent advances in oral squamous cell carcinoma. Semin. Cancer Biol. 2019;61:71–83. - PubMed
-
- Bonner J.A., et al. Radiotherapy plus cetuximab for locoregionally advanced head and neck cancer: 5-year survival data from a phase 3 randomised trial, and relation between cetuximab-induced rash and survival. Lancet Oncol. 2010;11(1):21–28. - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

