Tiny but mighty: Diverse functions of uORFs that regulate gene expression
- PMID: 39525088
- PMCID: PMC11550727
- DOI: 10.1016/j.csbj.2024.10.042
Tiny but mighty: Diverse functions of uORFs that regulate gene expression
Abstract
Upstream open reading frames (uORFs) are critical cis-acting regulators of downstream gene expression. Specifically, uORFs regulate translation by disrupting translation initiation or mediating mRNA decay. We herein summarize the effects of several uORFs that regulate gene expression in microbes to illustrate the detailed mechanisms mediating uORF functions. Microbes are ideal for uORF studies because of their prompt responses to stimuli. Recent studies revealed uORFs are ubiquitous in higher eukaryotes. Moreover, they influence various physiological processes in mammalian cells by regulating gene expression, mostly at the translational level. Research conducted using rapidly evolving methods for ribosome profiling combined with protein analyses and computational annotations showed that uORFs in mammalian cells control gene expression similar to microbial uORFs, but they also have unique tumorigenesis-related roles because of their protein-encoding capacities. We briefly introduce cutting-edge research findings regarding uORFs in mammalian cells.
Keywords: Gene expression and regulation; Microbes; Stress response; Transcription; Translation; UORFs.
© 2024 The Authors.
Conflict of interest statement
All the authors declare no conflict of interest.
Figures




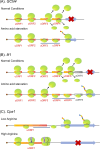

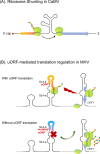
References
-
- Staib P., et al. Gene regulation and host adaptation mechanisms in Candida albicans. Int J Med Microbiol. 2001;291(2):183–188. - PubMed
-
- López-Maury L., Marguerat S., Bähler J. Tuning gene expression to changing environments: from rapid responses to evolutionary adaptation. Nat Rev Genet. 2008;9(8):583–593. - PubMed
-
- Des Marais D.L., Hernandez K.M., Juenger T.E. Genotype-by-environment interaction and plasticity: exploring genomic responses of plants to the abiotic environment. Annu Rev Ecol, Evol, Syst. 2013;44(44, 2013):5–29.
-
- Gao S., et al. Role of gene regulation and inter species interaction as a key factor in gut microbiota adaptation. Arch Microbiol. 2022;204(6):342. - PubMed
Publication types
LinkOut - more resources
Full Text Sources
