Exposure to live saprophytic Leptospira before challenge with a pathogenic serovar prevents severe leptospirosis and promotes kidney homeostasis
- PMID: 39527098
- PMCID: PMC11554301
- DOI: 10.7554/eLife.96470
Exposure to live saprophytic Leptospira before challenge with a pathogenic serovar prevents severe leptospirosis and promotes kidney homeostasis
Abstract
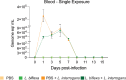
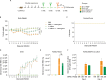
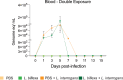
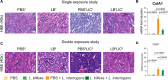
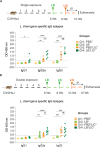
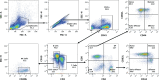
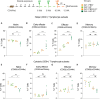
Previous studies demonstrated that Leptospira biflexa, a saprophytic species, triggers innate immune responses in the host during early infection. This raised the question of whether these responses could suppress a subsequent challenge with pathogenic Leptospira. We inoculated C3H/HeJ mice with a single or a double dose of L. biflexa before challenge with a pathogenic serovar, Leptospira interrogans serovar Copenhageni FioCruz (LIC). Pre-challenge exposure to L. biflexa did not prevent LIC dissemination and colonization of the kidney. However, it rescued weight loss and mouse survival thereby mitigating disease severity. Unexpectedly, there was correlation between rescue of overall health (weight gain, higher survival, lower kidney fibrosis marker ColA1) and higher shedding of LIC in urine. This stood in contrast to the L. biflexa unexposed LIC challenged control. Immune responses were dominated by increased frequency of effector T helper (CD4+) cells in spleen, as well as significant increases in serologic IgG2a. Our findings suggest that exposure to live saprophytic Leptospira primes the host to develop Th1 biased immune responses that prevent severe disease induced by a subsequent challenge with a pathogenic species. Thus, mice exposed to live saprophytic Leptospira before facing a pathogenic serovar may withstand infection with far better outcomes. Furthermore, a status of homeostasis may have been reached after kidney colonization that helps LIC complete its enzootic cycle.
Keywords: L. interrogans; Leptospira; commensalism; immunology; infectious disease; inflammation; kidney; leptospirosis; microbiology; mouse; vaccine.
© 2024, Kundu, Shetty et al.
Conflict of interest statement
SK, MG has a provisional patent application (application number 63/618,708) with the United States Patent and Trademark Office (USPTO), AS No competing interests declared
Figures



Update of
-
Exposure to live saprophytic Leptospira before challenge with a pathogenic serovar prevents severe leptospirosis and promotes kidney homeostasis.bioRxiv [Preprint]. 2024 Jul 1:2024.03.01.582981. doi: 10.1101/2024.03.01.582981. bioRxiv. 2024. Update in: Elife. 2024 Nov 11;13:RP96470. doi: 10.7554/eLife.96470. PMID: 38496604 Free PMC article. Updated. Preprint.
Similar articles
-
Exposure to live saprophytic Leptospira before challenge with a pathogenic serovar prevents severe leptospirosis and promotes kidney homeostasis.bioRxiv [Preprint]. 2024 Jul 1:2024.03.01.582981. doi: 10.1101/2024.03.01.582981. bioRxiv. 2024. Update in: Elife. 2024 Nov 11;13:RP96470. doi: 10.7554/eLife.96470. PMID: 38496604 Free PMC article. Updated. Preprint.
-
Mouse model for sublethal Leptospira interrogans infection.Infect Immun. 2015 Dec;83(12):4693-700. doi: 10.1128/IAI.01115-15. Epub 2015 Sep 28. Infect Immun. 2015. PMID: 26416909 Free PMC article.
-
The route of infection with Leptospira interrogans serovar Copenhageni affects the kinetics of bacterial dissemination and kidney colonization.PLoS Negl Trop Dis. 2020 Jan 6;14(1):e0007950. doi: 10.1371/journal.pntd.0007950. eCollection 2020 Jan. PLoS Negl Trop Dis. 2020. PMID: 31905198 Free PMC article.
-
Global proteome of the saprophytic strain Leptospira biflexa and comparative analysis with pathogenic strain Leptospira interrogans uncover new pathogenesis mechanisms.J Proteomics. 2024 Apr 15;297:105125. doi: 10.1016/j.jprot.2024.105125. Epub 2024 Feb 14. J Proteomics. 2024. PMID: 38364905
-
Anti-Leptospira immunoglobulin profiling in mice reveals strain specific IgG and persistent IgM responses associated with virulence and renal colonization.PLoS Negl Trop Dis. 2021 Mar 11;15(3):e0008970. doi: 10.1371/journal.pntd.0008970. eCollection 2021 Mar. PLoS Negl Trop Dis. 2021. PMID: 33705392 Free PMC article.
Cited by
-
TLR4 competence and mouse models of leptospirosis.bioRxiv [Preprint]. 2025 Feb 4:2025.02.03.636333. doi: 10.1101/2025.02.03.636333. bioRxiv. 2025. Update in: PLoS Negl Trop Dis. 2025 May 30;19(5):e0013163. doi: 10.1371/journal.pntd.0013163. PMID: 39975062 Free PMC article. Updated. Preprint.
-
Into the spotlight: A spatial study of potentially underreported leptospirosis among dengue-negative patients in São Paulo city, Brazil.PLoS Negl Trop Dis. 2025 Mar 5;19(3):e0012888. doi: 10.1371/journal.pntd.0012888. eCollection 2025 Mar. PLoS Negl Trop Dis. 2025. PMID: 40043023 Free PMC article.
-
Myocarditis and neutrophil-mediated vascular leakage but not cytokine storm associated with fatal murine leptospirosis.EBioMedicine. 2025 Feb;112:105571. doi: 10.1016/j.ebiom.2025.105571. Epub 2025 Jan 30. EBioMedicine. 2025. PMID: 39889371 Free PMC article.
-
Factors associated with the presence of anti-Leptospira spp. antibodies in persons experiencing homelessness in Brazil.Front Public Health. 2025 Jul 11;13:1596684. doi: 10.3389/fpubh.2025.1596684. eCollection 2025. Front Public Health. 2025. PMID: 40717952 Free PMC article.
-
Unveiling Leptospira prevalence and exposure in sanitation workers, a cross-sectional study in Ningbo City, China.Front Public Health. 2025 Jul 2;13:1627155. doi: 10.3389/fpubh.2025.1627155. eCollection 2025. Front Public Health. 2025. PMID: 40672915 Free PMC article.
References
-
- Barazzone GC, Teixeira AF, Azevedo BOP, Damiano DK, Oliveira MP, Nascimento A, Lopes APY. Revisiting the development of vaccines against pathogenic Leptospira: Innovative approaches, present challenges, and future perspectives. Frontiers in Immunology. 2021;12:760291. doi: 10.3389/fimmu.2021.760291. - DOI - PMC - PubMed
-
- Castiblanco-Valencia MM, Fraga TR, Breda LCD, Vasconcellos SA, Figueira CP, Picardeau M, Wunder E, Ko AI, Barbosa AS, Isaac L. Acquisition of negative complement regulators by the saprophyte Leptospira biflexa expressing LigA or LigB confers enhanced survival in human serum. Immunology Letters. 2016;173:61–68. doi: 10.1016/j.imlet.2016.03.005. - DOI - PMC - PubMed
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

