LSD1 deficiency in breast cancer cells promotes the formation of pre-metastatic niches
- PMID: 39528717
- PMCID: PMC11555121
- DOI: 10.1038/s41698-024-00751-2
LSD1 deficiency in breast cancer cells promotes the formation of pre-metastatic niches
Abstract
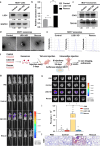
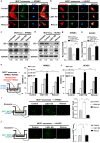
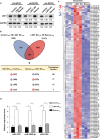
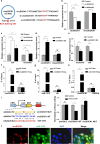
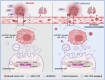
Lysine-specific demethylase 1 (LSD1), a histone demethylating enzyme, plays a crucial role in cancer metastasis. Studies show LSD1 knockout promotes breast cancer lung metastasis, but it's unknown if it alters the lung microenvironment for metastasis. In this study, we investigated the effects of exosomes from LSD1-knockdown (LSD1 KD) breast cancer cells on pre-metastatic niche formation. Injecting exosomes from LSD1 KD cells in mice resulted in a substantial increase in lung colonization by breast cancer cells, while treatment with exosomes derived from LSD1 KD cells decreased the expression of the ZO-1 and occludin, leading to increased vascular permeability. The LSD1 KD reduced the expression of circDOCK1, which augmented the levels of miR-1270 in exosomes. And miR-1270 inhibited ZO-1 expression in human endothelial cells, which enhanced their permeability. Our study uncovered a novel mechanism in which the LSD1 promotes the formation of pre-metastatic niches via the regulation of exosomal miRNA.
© 2024. The Author(s).
Conflict of interest statement
Figures



References
-
- Morgan, E. et al. The global landscape of esophageal squamous cell carcinoma and esophageal adenocarcinoma incidence and mortality in 2020 and projections to 2040: new estimates from GLOBOCAN 2020. Gastroenterology163, 649–658.e642 (2022). - PubMed
-
- Waks, A. G. & Winer, E. P. Breast cancer treatment: a review. JAMA321, 288–300 (2019). - PubMed
-
- Harbeck, N. et al. Breast cancer. Nat. Rev. Dis. Primers5, 66 (2019). - PubMed
-
- Nguyen, D. X., Bos, P. D. & Massague, J. Metastasis: from dissemination to organ-specific colonization. Nat. Rev. Cancer9, 274–284 (2009). - PubMed
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

