Contributions of mirror-image hair cell orientation to mouse otolith organ and zebrafish neuromast function
- PMID: 39531034
- PMCID: PMC11556791
- DOI: 10.7554/eLife.97674
Contributions of mirror-image hair cell orientation to mouse otolith organ and zebrafish neuromast function
Abstract
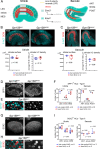
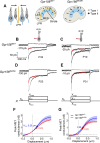
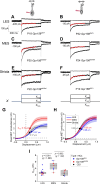
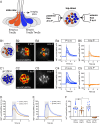
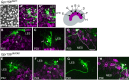
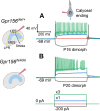
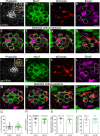
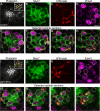
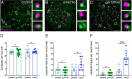
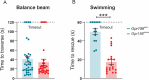
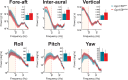
Otolith organs in the inner ear and neuromasts in the fish lateral-line harbor two populations of hair cells oriented to detect stimuli in opposing directions. The underlying mechanism is highly conserved: the transcription factor EMX2 is regionally expressed in just one hair cell population and acts through the receptor GPR156 to reverse cell orientation relative to the other population. In mouse and zebrafish, loss of Emx2 results in sensory organs that harbor only one hair cell orientation and are not innervated properly. In zebrafish, Emx2 also confers hair cells with reduced mechanosensory properties. Here, we leverage mouse and zebrafish models lacking GPR156 to determine how detecting stimuli of opposing directions serves vestibular function, and whether GPR156 has other roles besides orienting hair cells. We find that otolith organs in Gpr156 mouse mutants have normal zonal organization and normal type I-II hair cell distribution and mechano-electrical transduction properties. In contrast, gpr156 zebrafish mutants lack the smaller mechanically evoked signals that characterize Emx2-positive hair cells. Loss of GPR156 does not affect orientation-selectivity of afferents in mouse utricle or zebrafish neuromasts. Consistent with normal otolith organ anatomy and afferent selectivity, Gpr156 mutant mice do not show overt vestibular dysfunction. Instead, performance on two tests that engage otolith organs is significantly altered - swimming and off-vertical-axis rotation. We conclude that GPR156 relays hair cell orientation and transduction information downstream of EMX2, but not selectivity for direction-specific afferents. These results clarify how molecular mechanisms that confer bi-directionality to sensory organs contribute to function, from single hair cell physiology to animal behavior.
Keywords: afferent innervation; cell polarity; developmental biology; hair cells; inner ear; lateral line; mechano-electrical transduction; mouse; neuroscience; zebrafish.
Conflict of interest statement
KO, AJ, NH, AJ, HC, MD, RE, KC, KK, BT No competing interests declared
Figures


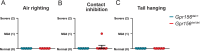
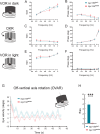

Update of
-
Contributions of mirror-image hair cell orientation to mouse otolith organ and zebrafish neuromast function.bioRxiv [Preprint]. 2024 Sep 6:2024.03.26.586740. doi: 10.1101/2024.03.26.586740. bioRxiv. 2024. Update in: Elife. 2024 Nov 12;13:RP97674. doi: 10.7554/eLife.97674. PMID: 39282410 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Molecular Biology Databases

