The ribosome-associated quality control pathway supports survival in the absence of non-stop ribosome rescue factors
- PMID: 39535229
- PMCID: PMC11633108
- DOI: 10.1128/mbio.02322-24
The ribosome-associated quality control pathway supports survival in the absence of non-stop ribosome rescue factors
Abstract
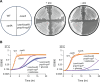
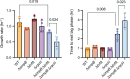
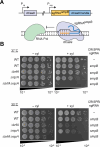
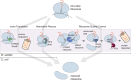
In bacteria, if a ribosome translates an mRNA lacking a stop codon it becomes stalled at the 3' end of the message. These ribosomes must be rescued by trans-translation or the alternative rescue factors (ArfA or ArfB). However, mounting evidence suggests that the ribosome quality control (RQC) pathway may also rescue non-stop ribosomes. Here, we surveyed the conservation of ribosome rescue pathways in >15,000 bacterial genomes. We found that trans-translation is conserved in >97% of bacterial genomes, while the other rescue pathways are restricted to particular phyla. We did not detect the gene encoding RqcH, the major mediator of RQC, in Proteobacteria (Pseudomonadota). In all Proteobacteria investigated to date, trans-translation is essential in the absence of the Arf proteins. Therefore, we tested whether expression of RQC components from Bacillus subtilis could rescue viability in the absence of trans-translation and ArfA in Escherichia coli. We found that the RQC pathway indeed functions in E. coli and rescues the well-documented synthetic lethal phenotype of ∆ssrA∆arfA. Moreover, we show that the RQC pathway in B. subtilis is essential in the absence of trans-translation and ArfA, further supporting a role for the RQC pathway in the rescue of non-stop ribosomes. Finally, we report a strong co-occurrence between RqcH and the ribosome splitting factor MutS2, but present experimental evidence that there are likely additional ribosome splitting factors beyond MutS2 in B. subtilis. Altogether, our work supports a role for RQC in non-stop ribosome rescue and provides a broad survey of ribosome rescue pathways in diverse bacteria.
Importance: In bacteria, it is estimated that 2%-4% of all translation reactions terminate with the ribosome stalled on a damaged mRNA lacking a stop codon. Mechanisms that rescue these ribosomes are essential for viability. We determined the functional overlap between the ribosome quality control pathway and the classical non-stop rescue systems [alternative rescue factor (ArfA) and trans-translation] in a representative Firmicute and Proteobacterium, phyla that are evolutionarily distinct. Furthermore, we used a bioinformatics approach to examine the conservation and overlap of various ribosome rescue systems in >15,000 species throughout the bacterial domain. These results provide key insights into ribosome rescue in diverse phyla.
Keywords: RNA binding proteins; bacteria; molecular genetics; ribosomes; trans-translation; translation.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Update of
-
RqcH supports survival in the absence of non-stop ribosome rescue factors.bioRxiv [Preprint]. 2024 Jul 12:2024.07.12.603306. doi: 10.1101/2024.07.12.603306. bioRxiv. 2024. Update in: mBio. 2024 Dec 11;15(12):e0232224. doi: 10.1128/mbio.02322-24. PMID: 39026760 Free PMC article. Updated. Preprint.
Similar articles
-
RqcH supports survival in the absence of non-stop ribosome rescue factors.bioRxiv [Preprint]. 2024 Jul 12:2024.07.12.603306. doi: 10.1101/2024.07.12.603306. bioRxiv. 2024. Update in: mBio. 2024 Dec 11;15(12):e0232224. doi: 10.1128/mbio.02322-24. PMID: 39026760 Free PMC article. Updated. Preprint.
-
A New Mechanism for Ribosome Rescue Can Recruit RF1 or RF2 to Nonstop Ribosomes.mBio. 2018 Dec 18;9(6):e02436-18. doi: 10.1128/mBio.02436-18. mBio. 2018. PMID: 30563899 Free PMC article.
-
trans-translation-mediated tight regulation of the expression of the alternative ribosome-rescue factor ArfA in Escherichia coli.Genes Genet Syst. 2011;86(3):151-63. doi: 10.1266/ggs.86.151. Genes Genet Syst. 2011. PMID: 21952205
-
Ribosome Rescue Pathways in Bacteria.Front Microbiol. 2021 Mar 18;12:652980. doi: 10.3389/fmicb.2021.652980. eCollection 2021. Front Microbiol. 2021. PMID: 33815344 Free PMC article. Review.
-
The fail-safe system to rescue the stalled ribosomes in Escherichia coli.Front Microbiol. 2014 Apr 10;5:156. doi: 10.3389/fmicb.2014.00156. eCollection 2014. Front Microbiol. 2014. PMID: 24782844 Free PMC article. Review.
Cited by
-
Molecular and structural basis of a subfamily of PrfH rescuing both the damaged and intact ribosomes stalled in translation.bioRxiv [Preprint]. 2025 Jan 9:2025.01.09.632186. doi: 10.1101/2025.01.09.632186. bioRxiv. 2025. PMID: 39829893 Free PMC article. Preprint.
-
IscR-tmRNA regulatory axis plays a key role in multiple stress response and pathogenicity in Aeromonas veronii.Virulence. 2025 Dec;16(1):2530659. doi: 10.1080/21505594.2025.2530659. Epub 2025 Jul 15. Virulence. 2025. PMID: 40665627 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

