Force-transducing molecular ensembles at growing microtubule tips control mitotic spindle size
- PMID: 39543105
- PMCID: PMC11564643
- DOI: 10.1038/s41467-024-54123-2
Force-transducing molecular ensembles at growing microtubule tips control mitotic spindle size
Abstract
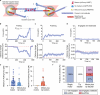
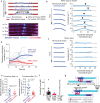
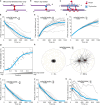
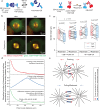
Correct mitotic spindle size is required for accurate chromosome segregation during cell division. It is controlled by mechanical forces generated by molecular motors and non-motor proteins acting on spindle microtubules. However, how forces generated by individual proteins enable bipolar spindle organization is not well understood. Here, we develop tools to measure contributions of individual molecules to this force balance. We show that microtubule plus-end binding proteins act at microtubule tips synergistically with minus-end directed motors to produce a system that can generate both pushing and pulling forces. To generate pushing force, the system harnesses forces generated by the growing tips of microtubules providing unique contribution to the force balance distinct from all other motors that act in the mitotic spindle. Our results reveal that microtubules are essential force generators for establishing spindle size and pave the way for understanding how mechanical forces can be fine-tuned to control the fidelity of chromosome segregation.
© 2024. The Author(s).
Conflict of interest statement
Figures

Similar articles
-
Kinesin-14 motors participate in a force balance at microtubule plus-ends to regulate dynamic instability.Proc Natl Acad Sci U S A. 2022 Feb 22;119(8):e2108046119. doi: 10.1073/pnas.2108046119. Proc Natl Acad Sci U S A. 2022. PMID: 35173049 Free PMC article.
-
The balance between KIFC3 and EG5 tetrameric kinesins controls the onset of mitotic spindle assembly.Nat Cell Biol. 2019 Sep;21(9):1138-1151. doi: 10.1038/s41556-019-0382-6. Epub 2019 Sep 2. Nat Cell Biol. 2019. PMID: 31481795
-
Antagonistic spindle motors and MAPs regulate metaphase spindle length and chromosome segregation.Curr Biol. 2013 Dec 2;23(23):2423-9. doi: 10.1016/j.cub.2013.10.023. Epub 2013 Nov 14. Curr Biol. 2013. PMID: 24239120 Free PMC article.
-
Molecular mechanisms of kinesin-14 motors in spindle assembly and chromosome segregation.J Cell Sci. 2017 Jul 1;130(13):2097-2110. doi: 10.1242/jcs.200261. J Cell Sci. 2017. PMID: 28668932 Review.
-
Assessment of Spindle Shape Control by Spindle Poleward Flux Measurements and FRAP Bulk Analysis.Methods Mol Biol. 2023;2604:113-125. doi: 10.1007/978-1-0716-2867-6_9. Methods Mol Biol. 2023. PMID: 36773229 Review.
Cited by
-
Stick-slip motion and universal statistics of cargo transport within living cells.bioRxiv [Preprint]. 2025 May 23:2025.05.19.654995. doi: 10.1101/2025.05.19.654995. bioRxiv. 2025. PMID: 40475663 Free PMC article. Preprint.
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

