Elucidating the pathobiology of Cerebellar Ataxia with Neuropathy and Vestibular Areflexia Syndrome (CANVAS) with its expanded RNA structure formation and proteinopathy
- PMID: 39543176
- PMCID: PMC11564700
- DOI: 10.1038/s41598-024-78947-6
Elucidating the pathobiology of Cerebellar Ataxia with Neuropathy and Vestibular Areflexia Syndrome (CANVAS) with its expanded RNA structure formation and proteinopathy
Abstract
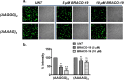
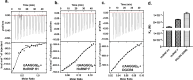
Numerous neurological disorders are linked to sequences rich in guanine repeats found in introns, exons, and regulatory regions of genes. These sequences have been observed to form stable G-quadruplex (GQ) structures both in vitro and in vitro. Cerebellar Ataxia with Neuropathy and Vestibular Areflexia Syndrome (CANVAS), a slowly progressive neurodegenerative disorder, is associated with the biallelic expansion of (AAGGG)n pathogenic repeats in the second intron of the RFC1 gene. Though these G-rich pathogenic repeats in other neurological diseases are associated with protein loss of function, RNA gain of function, and/or protein gain of function, not much is known about the pathological mechanism associated with CANVAS. Herein, we report the formation of stable GQ conformations in the CANVAS-associated repeats i.e., r(AAGGG)n, where 'r' stands for RNA. These GQs are critical regulators in neurological disorders leading to RNA foci formation and RNA binding protein sequestration. They also alter other causative processes like intron retention, which leads us to hypothesize a toxic Proteinopathy mechanism in CANVAS. Various biophysical and biomolecular assays characterized the interactions of three aggregation-prone RNA-binding proteins (RBPs): heterogeneous nuclear ribonucleoprotein H1/F (hnRNP H1/F), and DGCR8 with different pathogenic repeats [(AAGGG)9] in vitro, further affirming the hypothesis. The biophysical observations are further supported by molecular dynamics analysis and cell-based studies, putting us a step closer to elucidating the pathological mechanism(s) in CANVAS neuropathy, paving the way for the development of innovative therapeutic interventions.
Keywords: CANVAS; Proteinopathy; RFC1; RNA r(AAGGG)n; Repeat expansions.
© 2024. The Author(s).
Conflict of interest statement
Figures









References
-
- Szmulewicz, D. J. et al. Cerebellar ataxia, neuropathy, vestibular areflexia syndrome (CANVAS): a review of the clinical features and video-oculographic diagnosis. Ann. N Y Acad. Sci.1233, 139–147 (2011). - PubMed
-
- Montaut, S. et al. Biallelic RFC1-expansion in a French multicentric sporadic ataxia cohort. J. Neurol.268 (9), 3337–3343 (2021). - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources

