BRD4 Degradation Enhanced Glioma Sensitivity to Temozolomide by Regulating Notch1 via Glu-Modified GSH-Responsive Nanoparticles
- PMID: 39544152
- PMCID: PMC11672279
- DOI: 10.1002/advs.202409753
BRD4 Degradation Enhanced Glioma Sensitivity to Temozolomide by Regulating Notch1 via Glu-Modified GSH-Responsive Nanoparticles
Abstract
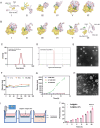
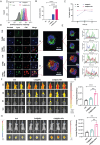
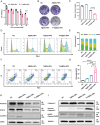
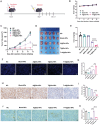
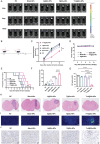
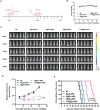
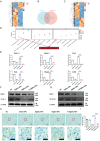
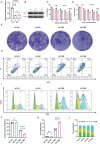
Temozolomide (TMZ) serves as the principal chemotherapeutic agent for glioma; nonetheless, its therapeutic efficacy is compromised by the rapid emergence of drug resistance, the inadequate targeting of glioma cells, and significant systemic toxicity. ARV-825 may play a role in modulating drug resistance by degrading the BRD4 protein, thereby exerting anti-glioma effects. Therefore, to surmount TMZ resistance and achieve efficient and specific drug delivery, a dual-targeted glutathione (GSH)-responsive nanoparticle system (T+A@Glu-NP) is designed and synthesized for the co-delivery of ARV-825 and TMZ. As anticipated, T+A@Glu-NPs significantly enhanced penetration of the blood-brain barrier (BBB), facilitated drug uptake by glioma cells, and exhibited efficient accumulation in brain tissue. Additionally, T+A@Glu-NPs exhibited augmented efficacy against glioma both in vitro and in vivo through the induction of apoptosis, inhibition of proliferation, and cell cycle arrest. Furthermore, mechanistic exploration revealed that T+A@Glu-NPs degraded the BRD4 protein, leading to the downregulation of Notch1 gene transcription and the inhibition of the Notch1 signaling pathway, thereby augmenting the therapeutic efficacy of glioma chemotherapy. Taken together, the findings suggest that T+A@Glu-NPs represents a novel and promising therapeutic strategy for glioma chemotherapy.
Keywords: ARV‐825; glioma chemotherapy, notch1; target delivery; temozolomide.
© 2024 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures


Similar articles
-
β-catenin contributes to cordycepin-induced MGMT inhibition and reduction of temozolomide resistance in glioma cells by increasing intracellular reactive oxygen species.Cancer Lett. 2018 Oct 28;435:66-79. doi: 10.1016/j.canlet.2018.07.040. Epub 2018 Aug 4. Cancer Lett. 2018. PMID: 30081068
-
Intranasal Delivery of Temozolomide-Conjugated Gold Nanoparticles Functionalized with Anti-EphA3 for Glioblastoma Targeting.Mol Pharm. 2021 Mar 1;18(3):915-927. doi: 10.1021/acs.molpharmaceut.0c00911. Epub 2021 Jan 8. Mol Pharm. 2021. PMID: 33417456
-
Nanoplatelets modified with RVG for targeted delivery of miR-375 and temozolomide to enhance gliomas therapy.J Nanobiotechnology. 2024 Oct 15;22(1):623. doi: 10.1186/s12951-024-02895-6. J Nanobiotechnology. 2024. PMID: 39402578 Free PMC article.
-
Fabrication of pH-responsive temozolomide (TMZ)-clacked tannic acid-altered zeolite imidazole nanoframeworks (ZIF-8) enhance anticancer activity and apoptosis induction in glioma cancer cells.J Biomater Sci Polym Ed. 2024 Sep;35(13):1978-1998. doi: 10.1080/09205063.2024.2364533. Epub 2024 Jul 2. J Biomater Sci Polym Ed. 2024. PMID: 38953298
-
Progress on the functions and mechanisms of natural products in anti-glioma therapy.Chin J Nat Med. 2025 May;23(5):541-559. doi: 10.1016/S1875-5364(25)60815-6. Chin J Nat Med. 2025. PMID: 40383611 Review.
Cited by
-
Overcoming temozolomide resistance in glioma: recent advances and mechanistic insights.Acta Neuropathol Commun. 2025 Jun 5;13(1):126. doi: 10.1186/s40478-025-02046-4. Acta Neuropathol Commun. 2025. PMID: 40468460 Free PMC article. Review.
References
-
- van den Bent M. J., Geurts M., French P. J., Smits M., Capper D., Bromberg J. E. C., Chang S. M., Lancet 2023, 402, 1564. - PubMed
-
- Alejo S., Palacios B. E., Venkata P. P., He Y., Li W., Johnson J. D., Chen Y., Jayamohan S., Pratap U. P., Clarke K., Zou Y., Lv Y., Weldon K., Viswanadhapalli S., Lai Z., Ye Z., Chen Y., Gilbert A. R., Suzuki T., Tekmal R. R., Zhao W., Zheng S., Vadlamudi R. K., Brenner A. J., Sareddy G. R., Neuro Oncol 2023, 25, 1249. - PMC - PubMed
-
- van den Bent M. J., Tesileanu C. M. S., Wick W., Sanson M., Brandes A. A., Clement P. M., Erridge S., Vogelbaum M. A., Nowak A. K., Baurain J. F., Mason W. P., Wheeler H., Chinot O. L., Gill S., Griffin M., Rogers L., Taal W., Rudà R., Weller M., McBain C., Reijneveld J., Enting R. H., Caparrotti F., Lesimple T., Clenton S., Gijtenbeek A., Lim E., Herrlinger U., Hau P., Dhermain F., et al., Lancet Oncol. 2021, 22, 813. - PMC - PubMed
MeSH terms
Substances
Grants and funding
- 32222046/National Natural Science Foundation of China
- 82172630/National Natural Science Foundation of China
- 2023NSFSC1931/Sichuan Science and Technology Program
- 2024YFFK0058/Sichuan Science and Technology Program
- 2024-YF05-00576-SN/Technology Innovation Research and Development Project of Chengdu Science and Technology Bureau
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
