A gentle introduction to pangenomics
- PMID: 39552065
- PMCID: PMC11570541
- DOI: 10.1093/bib/bbae588
A gentle introduction to pangenomics
Abstract
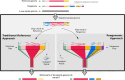
Pangenomes have emerged in response to limitations associated with traditional linear reference genomes. In contrast to a traditional reference that is (usually) assembled from a single individual, pangenomes aim to represent all of the genomic variation found in a group of organisms. The term 'pangenome' is currently used to describe multiple different types of genomic information, and limited language is available to differentiate between them. This is frustrating for researchers working in the field and confusing for researchers new to the field. Here, we provide an introduction to pangenomics relevant to both prokaryotic and eukaryotic organisms and propose a formalization of the language used to describe pangenomes (see the Glossary) to improve the specificity of discussion in the field.
Keywords: genomic variation; pangenome; presence–absence variation (PAV); reference bias.
© The Author(s) 2024. Published by Oxford University Press.
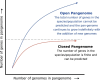
Figures




References
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources

