This is a preprint.
Combinatorial effector targeting (COMET) for transcriptional modulation and locus-specific biochemistry
- PMID: 39554033
- PMCID: PMC11565746
- DOI: 10.1101/2024.10.28.620517
Combinatorial effector targeting (COMET) for transcriptional modulation and locus-specific biochemistry
Abstract
Understanding how human gene expression is coordinately regulated by functional units of proteins across the genome remains a major biological goal. Here, we present COMET, a high-throughput screening platform for combinatorial effector targeting for the identification of transcriptional modulators. We generate libraries of combinatorial dCas9-based fusion proteins, containing two to six effector domains, allowing us to systematically investigate more than 110,000 combinations of effector proteins at endogenous human loci for their influence on transcription. Importantly, we keep full proteins or domains intact, maintaining catalytic cores and surfaces for protein-protein interactions. We observe more than 5800 significant hits that modulate transcription, we demonstrate cell type specific transcriptional modulation, and we further investigate epistatic relationships between our effector combinations. We validate unexpected combinations as synergistic or buffering, emphasizing COMET as both a method for transcriptional effector discovery, and as a functional genomics tool for identifying novel domain interactions and directing locus-specific biochemistry.
Conflict of interest statement
DECLARATION OF INTERESTS C.M.W., G.C.P., J.S.W., and L.A.G. have filed patent applications on the platform/materials presented in this report. J.S.W. serves as an advisor to and/or has equity in 5 AM Ventures, Amgen, Chroma Medicine, KSQ Therapeutics, Maze Therapeutics, Tenaya Therapeutics and Tessera Therapeutics. L.A.G. has filed patents on CRISPR tools and CRISPR functional genomics, is a co-founder of Chroma Medicine, and a consultant for Chroma Medicine. The other authors declare no competing interests.
Figures



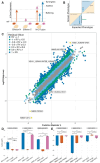

References
-
- Thakore P.I., D’Ippolito A.M., Song L., Safi A., Shivakumar N.K., Kabadi A.M., Reddy T.E., Crawford G.E., and Gersbach C.A. (2015). Highly specific epigenome editing by CRISPR-Cas9 repressors for silencing of distal regulatory elements. Nat. Methods 12, 1143–1149. 10.1038/nmeth.3630. - DOI - PMC - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
