Transcriptomics-Based Liquid Biopsy for Early Detection of Recurrence in Locally Advanced Gastric Cancer
- PMID: 39556695
- PMCID: PMC11653671
- DOI: 10.1002/advs.202406276
Transcriptomics-Based Liquid Biopsy for Early Detection of Recurrence in Locally Advanced Gastric Cancer
Abstract
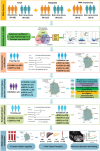
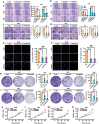
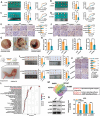
The study presents a transcriptomics-based liquid biopsy approach for early recurrence detection in locally advanced gastric cancer (LAGC). Four mRNA biomarkers (AGTR1, DNER, EPHA7, and SUSD5) linked to recurrence are identified through transcriptomic data analysis. A Risk Stratification Assessment (RSA) model combining these biomarkers with clinical features showed superior predictive accuracy for postoperative recurrence, with AUCs of 0.919 and 0.935 in surgical and liquid biopsy validation cohorts, respectively. Functional studies using human gastric cancer cell lines AGS and HGC-27 demonstrated that silencing the identified mRNA panel genes impaired cell migration, invasion, and proliferation. In vivo experiments further showed reduced tumor growth, metastasis, and lymphangiogenesis in mice, possibly mediated by the cAMP signaling pathway. This non-invasive approach offers significant potential for enhancing recurrence detection and enabling personalized treatment strategies, thereby improving patient outcomes in the management of LAGC.
Keywords: gastric cancer; liquid biopsy; mRNA Panel; recurrence detection; transcriptomics.
© 2024 The Author(s). Advanced Science published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflict of interest.
Figures





References
-
- Bray F., Laversanne M., Sung H., Ferlay J., Siegel R. L., Soerjomataram I., Jemal A., CA Cancer J Clin. 2024, 74, 229. - PubMed
-
- Siegel R. L., Giaquinto A. N., Jemal A., CA Cancer J Clin. 2024, 74, 12. - PubMed
-
- Smyth E. C., Nilsson M., Grabsch H. I., van Grieken N. C., Lordick F., Lancet. 2020, 396, 635. - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Molecular Biology Databases
Miscellaneous
