Genome evolution and diversity of wild and cultivated rice species
- PMID: 39557856
- PMCID: PMC11574199
- DOI: 10.1038/s41467-024-54427-3
Genome evolution and diversity of wild and cultivated rice species
Abstract
Wild species of crops serve as a valuable germplasm resource for breeding of modern cultivars. Rice (Oryza sativa L.) is a vital global staple food. However, research on genome evolution and diversity of wild rice species remains limited. Here, we present nearly complete genomes of 13 representative wild rice species. By integrating with four previously published genomes for pangenome analysis, a total of 101,723 gene families are identified across the genus, including 9834 (9.67%) core gene families. Additionally, 63,881 gene families absent in cultivated rice species but present in wild rice species are discovered. Extensive structural rearrangements, sub-genomes exchanges, widespread allelic variations, and regulatory sequence variations are observed in wild rice species. Interestingly, expanded but less diverse disease resistance genes in the genomes of cultivated rice, likely due to the loss of some resistance genes and the fixing and amplification of genes encoding resistance genes to specific diseases during domestication and artificial selection. This study not only reveals natural variations valuable for gene-level studies and breeding selection but also enhances our understanding on rice evolution and domestication.
© 2024. The Author(s).
Conflict of interest statement
Figures
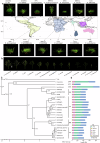





References
Publication types
MeSH terms
Associated data
Grants and funding
LinkOut - more resources
Full Text Sources

