Dominant negative variants in ITPR3 impair T cell Ca2+ dynamics causing combined immunodeficiency
- PMID: 39560673
- PMCID: PMC11577440
- DOI: 10.1084/jem.20220979
Dominant negative variants in ITPR3 impair T cell Ca2+ dynamics causing combined immunodeficiency
Abstract
The importance of calcium (Ca2+) as a second messenger in T cell signaling is exemplified by genetic deficiencies of STIM1 and ORAI1, which abolish store-operated Ca2+ entry (SOCE) resulting in combined immunodeficiency (CID). We report five unrelated patients with de novo missense variants in ITPR3, encoding a subunit of the inositol 1,4,5-trisphosphate receptor (IP3R), which forms a Ca2+ channel in the endoplasmic reticulum (ER) membrane responsible for the release of ER Ca2+ required to trigger SOCE, and for Ca2+ transfer to other organelles. The patients presented with CID, abnormal T cell Ca2+ homeostasis, incompletely penetrant ectodermal dysplasia, and multisystem disease. Their predominant T cell immunodeficiency is characterized by significant T cell lymphopenia, defects in late stages of thymic T cell development, and impaired function of peripheral T cells, including inadequate NF-κB- and NFAT-mediated, proliferative, and metabolic responses to activation. Pathogenicity is not due to haploinsufficiency, rather ITPR3 protein variants interfere with IP3R channel function leading to depletion of ER Ca2+ stores and blunted SOCE in T cells.
© 2024 Blanco et al.
Conflict of interest statement
Disclosures: S. Bahal reported grants from the Medical Research Council outside the submitted work. A.E. Handel reported grants from the Medical Research Council (UK), MyAware, and UCB-Pharma; and other from NIHR Oxford Health BRC outside the submitted work. S.O. Burns reported grants from CSL Behring and Pharming; other from CSL Behring, Glaxo Smith Klein, Baxalta US Inc., and Grifols; and personal fees from Biotest, Grifols, Pharming, and Takeda outside the submitted work. O. Gillham reported grants from Merck Sharp & Dohme (MSD) outside the submitted work. I. André reported grants from Smart Immune outside the submitted work. No other disclosures were reported.
Figures









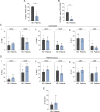
References
-
- Amrolia, P.J., Muccioli-Casadei G., Huls H., Adams S., Durett A., Gee A., Yvon E., Weiss H., Cobbold M., Gaspar H.B., et al. . 2006. Adoptive immunotherapy with allodepleted donor T-cells improves immune reconstitution after haploidentical stem cell transplantation. Blood. 108:1797–1808. 10.1182/blood-2006-02-001909 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

