InterPro: the protein sequence classification resource in 2025
- PMID: 39565202
- PMCID: PMC11701551
- DOI: 10.1093/nar/gkae1082
InterPro: the protein sequence classification resource in 2025
Abstract
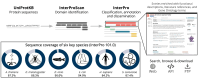
InterPro (https://www.ebi.ac.uk/interpro) is a freely accessible resource for the classification of protein sequences into families. It integrates predictive models, known as signatures, from multiple member databases to classify sequences into families and predict the presence of domains and significant sites. The InterPro database provides annotations for over 200 million sequences, ensuring extensive coverage of UniProtKB, the standard repository of protein sequences, and includes mappings to several other major resources, such as Gene Ontology (GO), Protein Data Bank in Europe (PDBe) and the AlphaFold Protein Structure Database. In this publication, we report on the status of InterPro (version 101.0), detailing new developments in the database, associated web interface and software. Notable updates include the increased integration of structures predicted by AlphaFold and the enhanced description of protein families using artificial intelligence. Over the past two years, more than 5000 new InterPro entries have been created. The InterPro website now offers access to 85 000 protein families and domains from its member databases and serves as a long-term archive for retired databases. InterPro data, software and tools are freely available.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures

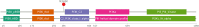
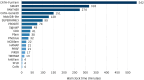
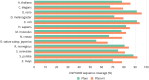
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Miscellaneous

