Examining the role of common variants in rare neurodevelopmental conditions
- PMID: 39567701
- PMCID: PMC11634775
- DOI: 10.1038/s41586-024-08217-y
Examining the role of common variants in rare neurodevelopmental conditions
Abstract
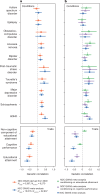
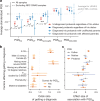
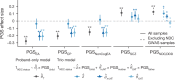
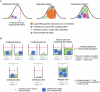
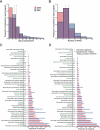
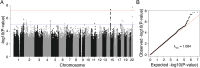
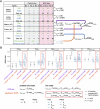
Although rare neurodevelopmental conditions have a large Mendelian component1, common genetic variants also contribute to risk2,3. However, little is known about how this polygenic risk is distributed among patients with these conditions and their parents nor its interplay with rare variants. It is also unclear whether polygenic background affects risk directly through alleles transmitted from parents to children, or whether indirect genetic effects mediated through the family environment4 also play a role. Here we addressed these questions using genetic data from 11,573 patients with rare neurodevelopmental conditions, 9,128 of their parents and 26,869 controls. Common variants explained around 10% of variance in risk. Patients with a monogenic diagnosis had significantly less polygenic risk than those without, supporting a liability threshold model5. A polygenic score for neurodevelopmental conditions showed only a direct genetic effect. By contrast, polygenic scores for educational attainment and cognitive performance showed no direct genetic effect, but the non-transmitted alleles in the parents were correlated with the child's risk, potentially due to indirect genetic effects and/or parental assortment for these traits4. Indeed, as expected under parental assortment, we show that common variant predisposition for neurodevelopmental conditions is correlated with the rare variant component of risk. These findings indicate that future studies should investigate the possible role and nature of indirect genetic effects on rare neurodevelopmental conditions, and consider the contribution of common and rare variants simultaneously when studying cognition-related phenotypes.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: M.E.H. is a cofounder of, consultant to and holds shares in Congenica, a genetics diagnostic company and is also a consultant to AstraZeneca Centre for Genomics Research.
Figures








References
-
- Kong, A. et al. The nature of nurture: effects of parental genotypes. Science359, 424–428 (2018). - PubMed
-
- Falconer, D. S. The inheritance of liability to certain diseases, estimated from the incidence among relatives. Ann. Hum. Genet.29, 51–76 (1965).
MeSH terms
Grants and funding
LinkOut - more resources
Full Text Sources
Medical

