Turnover of retroelements and satellite DNA drives centromere reorganization over short evolutionary timescales in Drosophila
- PMID: 39570997
- PMCID: PMC11620609
- DOI: 10.1371/journal.pbio.3002911
Turnover of retroelements and satellite DNA drives centromere reorganization over short evolutionary timescales in Drosophila
Abstract
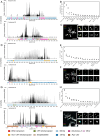
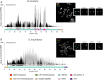
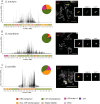
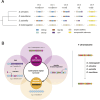
Centromeres reside in rapidly evolving, repeat-rich genomic regions, despite their essential function in chromosome segregation. Across organisms, centromeres are rich in selfish genetic elements such as transposable elements and satellite DNAs that can bias their transmission through meiosis. However, these elements still need to cooperate at some level and contribute to, or avoid interfering with, centromere function. To gain insight into the balance between conflict and cooperation at centromeric DNA, we take advantage of the close evolutionary relationships within the Drosophila simulans clade-D. simulans, D. sechellia, and D. mauritiana-and their relative, D. melanogaster. Using chromatin profiling combined with high-resolution fluorescence in situ hybridization on stretched chromatin fibers, we characterize all centromeres across these species. We discovered dramatic centromere reorganization involving recurrent shifts between retroelements and satellite DNAs over short evolutionary timescales. We also reveal the recent origin (<240 Kya) of telocentric chromosomes in D. sechellia, where the X and fourth centromeres now sit on telomere-specific retroelements. Finally, the Y chromosome centromeres, which are the only chromosomes that do not experience female meiosis, do not show dynamic cycling between satDNA and TEs. The patterns of rapid centromere turnover in these species are consistent with genetic conflicts in the female germline and have implications for centromeric DNA function and karyotype evolution. Regardless of the evolutionary forces driving this turnover, the rapid reorganization of centromeric sequences over short evolutionary timescales highlights their potential as hotspots for evolutionary innovation.
Copyright: © 2024 Courret et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
Islands of retroelements are major components of Drosophila centromeres.PLoS Biol. 2019 May 14;17(5):e3000241. doi: 10.1371/journal.pbio.3000241. eCollection 2019 May. PLoS Biol. 2019. PMID: 31086362 Free PMC article.
-
Comparative Analyses of Gibbon Centromeres Reveal Dynamic Genus-Specific Shifts in Repeat Composition.Mol Biol Evol. 2021 Aug 23;38(9):3972-3992. doi: 10.1093/molbev/msab148. Mol Biol Evol. 2021. PMID: 33983366 Free PMC article.
-
Dynamic Evolution of Euchromatic Satellites on the X Chromosome in Drosophila melanogaster and the simulans Clade.Mol Biol Evol. 2020 Aug 1;37(8):2241-2256. doi: 10.1093/molbev/msaa078. Mol Biol Evol. 2020. PMID: 32191304 Free PMC article.
-
Transposable elements: genome innovation, chromosome diversity, and centromere conflict.Chromosome Res. 2018 Mar;26(1-2):5-23. doi: 10.1007/s10577-017-9569-5. Epub 2018 Jan 13. Chromosome Res. 2018. PMID: 29332159 Free PMC article. Review.
-
What makes a centromere?Exp Cell Res. 2020 Apr 15;389(2):111895. doi: 10.1016/j.yexcr.2020.111895. Epub 2020 Feb 6. Exp Cell Res. 2020. PMID: 32035948 Review.
Cited by
-
Centromeres drive and take a break.Chromosome Res. 2025 Aug 4;33(1):17. doi: 10.1007/s10577-025-09777-z. Chromosome Res. 2025. PMID: 40759764 Free PMC article. Review.
-
Analysis of 30 chromosome-level Drosophila genome assemblies reveals dynamic evolution of centromeric satellite repeats.Genome Biol. 2025 Mar 18;26(1):63. doi: 10.1186/s13059-025-03527-4. Genome Biol. 2025. PMID: 40102968 Free PMC article.
-
Controlling and controlled elements: highlights of the year in mobile DNA research.Mob DNA. 2024 Dec 31;15(1):27. doi: 10.1186/s13100-024-00340-x. Mob DNA. 2024. PMID: 39741304 Free PMC article. No abstract available.
-
Patterns of crossover distribution in Drosophila mauritiana necessitate a re-thinking of the centromere effect on crossing over.Genetics. 2025 May 8;230(1):iyaf039. doi: 10.1093/genetics/iyaf039. Genetics. 2025. PMID: 40052765 Free PMC article.
-
Chromosome-scale scaffolds of the fungus gnat genome reveal multi-Mb-scale chromosome-folding interactions, centromeric enrichments of retrotransposons, and candidate telomere sequences.BMC Genomics. 2025 May 5;26(1):443. doi: 10.1186/s12864-025-11573-2. BMC Genomics. 2025. PMID: 40325439 Free PMC article.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases

