DUSP6 regulates Notch1 signalling in colorectal cancer
- PMID: 39572549
- PMCID: PMC11582695
- DOI: 10.1038/s41467-024-54383-y
DUSP6 regulates Notch1 signalling in colorectal cancer
Abstract
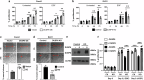
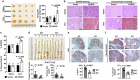
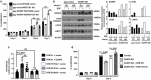
Notch1 plays various roles in cancer development, and Notch1-induced transactivation is controlled by phosphorylation of its cleaved intracellular domain. However, it is unclear whether there are phosphatases capable of dephosphorylating the cleaved Notch1 transmembrane/intracellular region (NTM) to regulate its function. Here, we show that DUSP6 can function as a phosphatase for Notch1, thereby regulating NTM stability and transcriptional activity, thus influencing colorectal cancer (CRC) development. In human CRC cells, elevated DUSP6 expression correlates with increased NTM levels, leading to enhanced CRC cell proliferation both in vitro and in vivo. High tumoral DUSP6 protein expression is associated with poorer overall CRC patient survival. In mice, DUSP6 deficiency results in reduced CRC development. Mechanistically, DUSP6 dephosphorylates phospho-Y2116, which in turn reduces NTM ubiquitination, leading to increased NTM stability and transcriptional activity. As a result, the expression of Notch1-targeted proliferation genes is increased to promote tumour cell growth.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures




References
-
- de la Cruz-Morcillo, M. A. et al. P38MAPK is a major determinant of the balance between apoptosis and autophagy triggered by 5-fluorouracil: implication in resistance. Oncogene31, 1073–1085 (2012). - PubMed
-
- Karapetis, C. S. et al. K-ras mutations and benefit from cetuximab in advanced colorectal cancer. N. Engl. J. Med.359, 1757–1765 (2008). - PubMed
-
- Camps, M., Nichols, A. & Arkinstall, S. Dual specificity phosphatases: a gene family for control of MAP kinase function. FASEB J.14, 6–16 (2000). - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials
Miscellaneous

