MicroRNA-145-5p inhibits the tumorigenesis of breast cancer through SENP2-regulated ubiquitination of ERK2
- PMID: 39578257
- PMCID: PMC11584840
- DOI: 10.1007/s00018-024-05505-8
MicroRNA-145-5p inhibits the tumorigenesis of breast cancer through SENP2-regulated ubiquitination of ERK2
Abstract
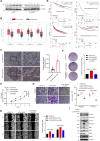
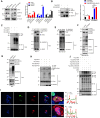
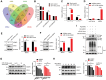
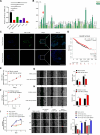
Breast carcinoma exhibits the highest incidence among various cancers and is the foremost cause of mortality in women. Increasing evidence shows that SUMOylation of proteins plays a critical role in the progression of breast cancer; however, the role of SENP2 and its molecular mechanism in breast cancer remain underexplored. Here, we discerned that SENP2 promoted the tumorigenesis of breast cancer both in vitro and in vivo. Furthermore, we identified that ERK2 was SUMOylated and that SENP2 played a role by deconjugating ERK2 SUMOylation in breast cancer. SUMOylation of ERK2 promoted its ubiquitin-proteasomal degradation, thus inhibiting the epithelial-to-mesenchymal transition in breast cancer cells. Furthermore, microRNA-145-5p (miR-145-5p) has emerged as a scarce commodity in breast cancer and binds to the 3'-untranslated region of SENP2 mRNA to govern the regulatory dynamics of SENP2 expression. Finally, miR-145-5p inhibits SENP2 transcription, enhances ERK2 SUMOylation, and ultimately suppresses the progression of breast cancer. These revelations suggest evolving ideas for the miR-145-5p-SENP2 axis in therapeutic intervention, thus heralding transformative prospects for the clinical management of breast cancer.
Keywords: Breast cancer; ERK2; MicroRNA-145-5p; SENP2; SUMOylation.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: All animal studies were performed according to the protocol “Guide for the Care and Use of Laboratory Animals”, which was approved by the esteemed Institutional Animal Care and Use Committee of Shaanxi Normal University (Xi’an, China, 202412004), and all manipulations were carried out according to established guidelines. All human specimens were analyzed for the current study with appropriate IRB approved by the Affiliated Hospital of Southwest Medical University (Luzhou, China, L2021012), and the studies abide by the Declaration of Helsinki principles. Consent for publication: All the authors involved in this manuscript give the consent for the publication. Conflict of interest: The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures



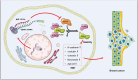
References
-
- Chen W et al (2016) Cancer statistics in China, 2015. CA Cancer J Clin 66(2):115–132 - PubMed
-
- Siegel RL, Miller KD, Jemal A (2019) Cancer statistics, 2019. CA Cancer J Clin 69(1):7–34 - PubMed
-
- Harbeck N, Gnant M (2017) Breast cancer Lancet 389(10074):1134–1150 - PubMed
-
- Dent R et al (2007) Triple-negative breast cancer: clinical features and patterns of recurrence. Clin Cancer Res 13(15 Pt 1):4429–4434 - PubMed
MeSH terms
Substances
Grants and funding
- 82273395/National Natural Science Foundation of China
- 81870241/National Natural Science Foundation of China
- 82203387/National Natural Science Foundation of China
- GK202205009/Fundamental Research Funds for the Central Universities
- GK202202006/Fundamental Research Funds for the Central Universities
- GK202207003/Fundamental Research Funds for the Central Universities
- LHRCTS23089/Excellent Graduate Training Program of Shaanxi Normal University
- 2023BSHYDZZ117/Shaanxi Province Postdoctoral Science Foundation
- AM2022017/Macao Youth Scholars Program
- LHRCCX23174/Excellent Graduate Training Program of Shaanxi Normal University
LinkOut - more resources
Full Text Sources
Medical
Miscellaneous

