miRTarBase 2025: updates to the collection of experimentally validated microRNA-target interactions
- PMID: 39578692
- PMCID: PMC11701613
- DOI: 10.1093/nar/gkae1072
miRTarBase 2025: updates to the collection of experimentally validated microRNA-target interactions
Abstract
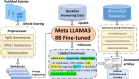
MicroRNAs (miRNAs) are small non-coding RNAs (18-26 nucleotides) that regulate gene expression by interacting with target mRNAs, affecting various physiological and pathological processes. miRTarBase, a database of experimentally validated miRNA-target interactions (MTIs), now features over 3 817 550 validated MTIs from 13 690 articles, significantly expanding its previous version. The updated database includes miRNA interactions with therapeutic agents, revealing roles in drug resistance and therapeutic strategies. It also highlights miRNAs as predictive, safety and monitoring biomarkers for toxicity assessment, clinical treatment guidance and therapeutic optimization. The expansion of miRNA-mRNA and miRNA-miRNA networks allows the identification of key regulatory genes and co-regulatory miRNAs, providing deeper insights into miRNA functions and critical target genes. Information on oxidized miRNA sequences has been added, shedding light on how oxidative modifications influence miRNA targeting and regulation. The integration of the LLAMA3 model into the NLP pipeline, alongside prompt engineering, enables the efficient identification of MTIs and miRNA-disease associations without large training datasets. An updated data integration and a redesigned user interface enhance accessibility, reinforcing miRTarBase as an essential resource for molecular oncology, drug development and related fields. The updated miRTarBase is available at https://mirtarbase.cuhk.edu.cn/∼miRTarBase/miRTarBase_2025.
© The Author(s) 2024. Published by Oxford University Press on behalf of Nucleic Acids Research.
Figures



References
MeSH terms
Substances
Grants and funding
- JCYJ20220530143615035/Shenzhen Science and Technology Program
- 32070674/National Natural Science Foundation of China
- 2024A0505050001/Guangdong S&T programme
- LGKCSDPT2024001/Warshel Institute for Computational Biology funding from Shenzhen City and Longgang District
- HZQB-KCZYB-2020056/Shenzhen-Hong Kong Cooperation Zone for Technology and Innovation
LinkOut - more resources
Full Text Sources

