Recent Advances in Labeling-Based Quantitative Glycomics: From High-Throughput Quantification to Structural Elucidation
- PMID: 39580675
- PMCID: PMC11735667
- DOI: 10.1002/pmic.202400057
Recent Advances in Labeling-Based Quantitative Glycomics: From High-Throughput Quantification to Structural Elucidation
Abstract
Glycosylation, a crucial posttranslational modification (PTM), plays important roles in numerous biological processes and is linked to various diseases. Despite its significance, the structural complexity and diversity of glycans present significant challenges for mass spectrometry (MS)-based quantitative analysis. This review aims to provide an in-depth overview of recent advancements in labeling strategies for N-glycomics and O-glycomics, with a specific focus on enhancing the sensitivity, specificity, and throughput of MS analyses. We categorize these advancements into three major areas: (1) the development of isotopic/isobaric labeling techniques that significantly improve multiplexing capacity and throughput for glycan quantification; (2) novel methods that aid in the structural elucidation of complex glycans, particularly sialylated and fucosylated glycans; and (3) labeling techniques that enhance detection ionization efficiency, separation, and sensitivity for matrix-assisted laser desorption/ionization (MALDI)-MS and capillary electrophoresis (CE)-based glycan analysis. In addition, we highlight emerging trends in single-cell glycomics and bioinformatics tools that have the potential to revolutionize glycan quantification. These developments not only expand our understanding of glycan structures and functions but also open new avenues for biomarker discovery and therapeutic applications. Through detailed discussions of methodological advancements, this review underscores the critical role of derivatization methods in advancing glycan identification and quantification.
Keywords: N‐glycan; O‐glycan; derivatization; glycan structure elucidation; sialylated glycans.
© 2024 The Author(s). Proteomics published by Wiley‐VCH GmbH.
Conflict of interest statement
The authors declare no conflicts of interest.
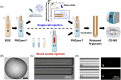
Figures




Similar articles
-
Recent advances in analytical methods and bioinformatic tools for quantitative glycomics.Anal Bioanal Chem. 2025 Apr;417(10):1947-1959. doi: 10.1007/s00216-025-05778-3. Epub 2025 Feb 13. Anal Bioanal Chem. 2025. PMID: 39948299 Review.
-
Quantitative O-glycomics based on improvement of the one-pot method for nonreductive O-glycan release and simultaneous stable isotope labeling with 1-(d0/d5)phenyl-3-methyl-5-pyrazolone followed by mass spectrometric analysis.J Proteomics. 2017 Jan 6;150:18-30. doi: 10.1016/j.jprot.2016.08.012. Epub 2016 Aug 29. J Proteomics. 2017. PMID: 27585995
-
A MALDI-MS-based quantitative targeted glycomics (MALDI-QTaG) for total N-glycan analysis.Biotechnol Lett. 2015 Oct;37(10):2019-25. doi: 10.1007/s10529-015-1881-6. Epub 2015 Jun 11. Biotechnol Lett. 2015. PMID: 26063621
-
Hitting the sweet spot with capillary electrophoresis: advances in N-glycomics and glycoproteomics.Curr Opin Biotechnol. 2021 Oct;71:182-190. doi: 10.1016/j.copbio.2021.07.013. Epub 2021 Aug 23. Curr Opin Biotechnol. 2021. PMID: 34438131 Review.
-
A novel, simplified strategy of relative quantification N-glycan: Quantitative glycomics using electrospray ionization mass spectrometry through the stable isotopic labeling by transglycosylation reaction of mutant enzyme Endo-M-N175Q.J Pharm Biomed Anal. 2018 Feb 5;149:365-373. doi: 10.1016/j.jpba.2017.11.032. Epub 2017 Nov 12. J Pharm Biomed Anal. 2018. PMID: 29145098
Cited by
-
Recent advances in analytical methods and bioinformatic tools for quantitative glycomics.Anal Bioanal Chem. 2025 Apr;417(10):1947-1959. doi: 10.1007/s00216-025-05778-3. Epub 2025 Feb 13. Anal Bioanal Chem. 2025. PMID: 39948299 Review.
-
A comprehensive review on computational metabolomics: Advancing multiscale analysis through in-silico approaches.Comput Struct Biotechnol J. 2025 Jul 13;27:3191-3215. doi: 10.1016/j.csbj.2025.07.016. eCollection 2025. Comput Struct Biotechnol J. 2025. PMID: 40735430 Free PMC article. Review.
References
Publication types
MeSH terms
Substances
Grants and funding
- S10 OD025084/OD/NIH HHS/United States
- S10OD028473/NIH Office of the Director
- S10OD025084/NIH Office of the Director
- P41GM108538/GM/NIGMS NIH HHS/United States
- S10 RR029531/RR/NCRR NIH HHS/United States
- P41 GM108538/GM/NIGMS NIH HHS/United States
- R01AG078794/AG/NIA NIH HHS/United States
- R01 AG078794/AG/NIA NIH HHS/United States
- S10RR029531/RR/NCRR NIH HHS/United States
- S10 OD028473/OD/NIH HHS/United States
- R01 AG052324/AG/NIA NIH HHS/United States
- R01 DK071801/DK/NIDDK NIH HHS/United States
- R01AG052324/AG/NIA NIH HHS/United States
LinkOut - more resources
Full Text Sources

