Serological evidence of sarbecovirus exposure along Sunda pangolin trafficking pathways
- PMID: 39593133
- PMCID: PMC11600613
- DOI: 10.1186/s12915-024-02074-x
Serological evidence of sarbecovirus exposure along Sunda pangolin trafficking pathways
Abstract
Background: Early in the coronavirus disease 2019 (COVID-19) pandemic, Sunda pangolins (Manis javanica) involved in the illegal wildlife trade in mainland China were identified as hosts of severe acute respiratory syndrome-related coronaviruses (SARSr-CoVs). Although it is unconfirmed whether pangolins or other traded wildlife served as intermediate hosts for severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2), the trafficking of pangolins presents a clear risk for transmission of viruses with zoonotic and epizootic potential regardless. We have investigated the origins of pangolin carcasses seized in Hong Kong and have evaluated their potential exposure to SARSr-CoVs, other coronaviruses, and paramyxoviruses, aiming to address a gap in our knowledge with regard to the role of wildlife trade in the maintenance and emergence of pathogens with zoonotic and epizootic potential.
Results: Using a combination of virological and wildlife forensics tools, we investigated 89 Sunda pangolin carcasses seized by Hong Kong authorities during anti-smuggling operations in the territory conducted in 2013 (n = 1) and 2018 (n = 88). Swabs, organ tissues, blood, and other body fluids were collected during post-mortem examination. Two enzyme-linked immunosorbent assays (ELISAs), which employ a double-antigen sandwich format, were used to detect antibodies reactive against SARSr-CoVs. One individual was found to be seropositive with support from both methods, while five individuals exhibited a putatively seropositive result from one ELISA method. Polymerase chain reaction (PCR) screening for coronavirus and paramyxovirus ribonucleic acid (RNA) did not yield any positives. Based on genomic data, the seropositive individual was determined to have likely originated from Java, while the putatively seropositive individuals were determined to have originated from populations in Borneo, Java, and Singapore/Sumatra.
Conclusions: While the role of pangolins in the evolution and ecology of SARS-CoV-2 is uncertain, our results suggest susceptibility and potential exposure of pangolins to SARSr-CoVs, occurring naturally or associated with the illegal trafficking of these animals. Complex dynamics between natural populations, traded individuals, and pathogen susceptibility complicate conclusions about the role of pangolins, as well as other host species, in the ecology of SARSr-CoVs and potentially zoonotic viruses with risk of future emergence.
Keywords: Conservation forensics; Coronavirus; ELISA; One Health; Pangolins; Paramyxovirus; Population genomics; SARS-related virus; Sarbecovirus; Serology.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Not applicable. Consent for publication: Not applicable. Competing interests: The authors declare that they have no competing interests.
Figures
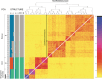
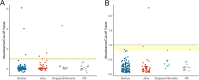

References
MeSH terms
Substances
Grants and funding
- R7021-20/Research Impact Fund
- COVID190223/Health and Medical Research Fund
- 31922087/National Natural Science Foundation of China's Excellent Young Scientist Fund
- 2019B121205009/Guangdong-Hong Kong-Macau Joint Laboratory Program
- HZQB-KCYZ-2021014/Shenzhen-Hong Kong Science and Technology Innovation Program
LinkOut - more resources
Full Text Sources
Miscellaneous

