Cancer-Associated Fibroblast-Derived FGF7 Promotes Clear Cell Renal Cell Carcinoma Progression and Macrophage Infiltration
- PMID: 39594574
- PMCID: PMC11593278
- DOI: 10.3390/cells13221824
Cancer-Associated Fibroblast-Derived FGF7 Promotes Clear Cell Renal Cell Carcinoma Progression and Macrophage Infiltration
Abstract
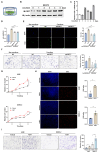
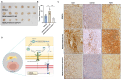
As the predominant stromal cells in the ccRCC surrounding environment, cancer-associated fibroblasts (CAFs) have been established as supportive of tumor growth. However, the detailed molecular mechanisms underlying the supporting role of CAFs in ccRCC have not been well characterized. Evidence from the clustering consensus analysis, single-cell analysis, and the experimental results illustrate that CAF-derived FGF7 plays a crucial role as a signaling mediator between CAFs and ccRCC tumor cells. Mechanistically, CAF-derived FGF7 triggers AKT activation to promote cell growth and cell invasion of ccRCC tumor cells. As a response, ccRCC tumor cells stimulate STAT3-mediated transcriptional regulation, directly increasing FGF7 expression at the chromatin level in CAFs. Moreover, there exists a positive clinical correlation between the abundance of CAFs, FGF7 expression, and the infiltration of M2 type macrophages. The RENCA in vivo mouse model also confirmed that FGF7 depletion could impede RCC development by reducing the recruitment of M2 type macrophages. Overall, this study delineates a key signaling axis governing the crosstalk between CAFs and ccRCC tumor cells, highlighting FGF7 as a promising therapeutic target of ccRCC.
Keywords: CAFs; FGF7; ccRCC.
Conflict of interest statement
No competing interests will be declared.
Figures




References
-
- Marona P., Górka J., Kwapisz O., Jura J., Rys J., Hoffman R.M., Miekus K. Resistance to tyrosine kinase inhibitors promotes renal cancer progression through MCPIP1 tumor-suppressor downregulation and c-Met activation. Cell Death Dis. 2022;13:814. doi: 10.1038/s41419-022-05251-4. - DOI - PMC - PubMed
-
- Msaouel P., Goswami S., Thall P.F., Wang X., Yuan Y., Jonasch E., Gao J., Campbell M.T., Shah A.Y., Corn P.G., et al. A phase 1-2 trial of sitravatinib and nivolumab in clear cell renal cell carcinoma following progression on antiangiogenic therapy. Sci. Transl. Med. 2022;14:eabm6420. doi: 10.1126/scitranslmed.abm6420. - DOI - PMC - PubMed
Publication types
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Other Literature Sources
Medical
Research Materials
Miscellaneous

