TMEM206 Contributes to Cancer Hallmark Functions in Colorectal Cancer Cells and Is Regulated by p53 in a p21-Dependent Manner
- PMID: 39594575
- PMCID: PMC11593115
- DOI: 10.3390/cells13221825
TMEM206 Contributes to Cancer Hallmark Functions in Colorectal Cancer Cells and Is Regulated by p53 in a p21-Dependent Manner
Abstract
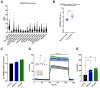
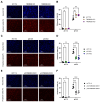
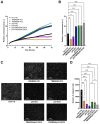
Acid-induced ion flux plays a role in pathologies where tissue acidification is prevalent, including cancer. In 2019, TMEM206 was identified as the molecular component of acid-induced chloride flux. Localizing to the plasma membrane, TMEM206 contributes to cellular processes like acid-induced cell death. Since over 50% of human cancers carry loss of function mutations in the p53 gene, we aimed to analyze how TMEM206 is regulated by p53 and its role in cancer hallmark function and acid-induced cell death in HCT116 colorectal cancer (CRC) cells. We generated p53-deficient HCT116 cells and assessed TMEM206-mediated Cl- currents and transcriptional regulation using the patch-clamp and a dual-luciferase reporter assay, respectively. To investigate the contribution of TMEM206 to cancer hallmark functions, we performed migration and metabolic activity assays. The role of TMEM206 in p53-mediated acid-induced cell death was assessed with cell death assays. The TMEM206 mRNA level was significantly elevated in human primary CRC tumors. TMEM206 knockout increased acid-induced cell death and reduced proliferation and migration, indicating a role for TMEM206 in these cancer hallmark functions. Furthermore, we observed increased TMEM206 mRNA levels and currents in HCT116 p53 knockout cells. This phenotype can be rescued by transient overexpression of p53 but not by overexpression of dysfunctional p53. In addition, our data suggest that TMEM206 may mediate cancer hallmark functions within p53-associated pathways. TMEM206 promoter activity is not altered by p53 overexpression. Conversely, knockout of p21, a major target gene of p53, increased TMEM206-mediated currents, suggesting expression control of TMEM206 by p21 downstream signaling. Our results show that in colorectal cancer cells, TMEM206 expression is elevated, contributes to cancer hallmark functions, and its regulation is dependent on p53 through a p21-dependent mechanism.
Keywords: TMEM206; acid-induced cell death; colorectal cancer; ion channel; p21; p53.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures


References
-
- Bell D.C., Leanza L., Gentile S., Sauter D.R. News and Views on Ion Channels in Cancer: Is Cancer a Channelopathy? Front. Pharmacol. 2023;14:1258933. doi: 10.3389/fphar.2023.1258933. - DOI
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

