Analysis of a cell-free DNA-based cancer screening cohort links fragmentomic profiles, nuclease levels, and plasma DNA concentrations
- PMID: 39603706
- PMCID: PMC11789642
- DOI: 10.1101/gr.279667.124
Analysis of a cell-free DNA-based cancer screening cohort links fragmentomic profiles, nuclease levels, and plasma DNA concentrations
Abstract
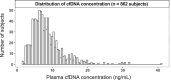
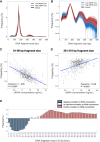
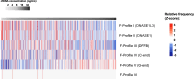
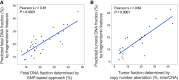
The concentration of circulating cell-free DNA (cfDNA) in plasma is an important determinant of the robustness of liquid biopsies. However, biological mechanisms that lead to inter-individual differences in cfDNA concentrations remain unexplored. The concentration of plasma cfDNA is governed by an interplay between its release and clearance. We hypothesized that cfDNA clearance by nucleases might be one mechanism that contributes toward inter-individual variations in cfDNA concentrations. We performed fragmentomic analysis of the plasma cfDNA from 862 healthy individuals, with a cfDNA concentration range of 1.61-41.01 ng/mL. We observed an increase in large DNA fragments (231-600 bp), a decreased frequencies of shorter DNA fragments (20-160 bp), and an increased frequency of G-end motifs with increasing cfDNA concentrations. End motif deconvolution analysis revealed a decreased contribution of DNASE1L3 and DFFB in subjects with higher cfDNA concentration. The five subjects with the highest plasma DNA concentration (top 0.58%) had aberrantly decreased levels of DNASE1L3 protein in plasma. The cfDNA concentration could be inferred from the fragmentomic profile through machine learning and was well correlated to the measured cfDNA concentration. Such an approach could infer the fractional DNA concentration from particular tissue types, such as the fetal and tumor fraction. This work shows that individuals with different cfDNA concentrations are associated with characteristic fragmentomic patterns of the cfDNA pool and that nuclease-mediated clearance of DNA is a key parameter that affects cfDNA concentration. Understanding these mechanisms has facilitated the enhanced measurement of cfDNA species of clinical interest, including circulating fetal and tumor DNA.
© 2025 Malki et al.; Published by Cold Spring Harbor Laboratory Press.
Figures


References
-
- Alborelli I, Generali D, Jermann P, Cappelletti MR, Ferrero G, Scaggiante B, Bortul M, Zanconati F, Nicolet S, Haegele J, et al. 2019. Cell-free DNA analysis in healthy individuals by next-generation sequencing: a proof of concept and technical validation study. Cell Death Dis 10: 534. 10.1038/s41419-019-1770-3 - DOI - PMC - PubMed
-
- Chan RWY, Serpas L, Ni M, Volpi S, Hiraki LT, Tam LS, Rashidfarrokhi A, Wong PCH, Tam LHP, Wang Y, et al. 2020. Plasma DNA profile associated with DNASE1L3 gene mutations: clinical observations, relationships to nuclease substrate preference, and in vivo correction. Am J Hum Genet 107: 882–894. 10.1016/j.ajhg.2020.09.006 - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
