A proteome-wide quantitative platform for nanoscale spatially resolved extraction of membrane proteins into native nanodiscs
- PMID: 39609567
- PMCID: PMC11810782
- DOI: 10.1038/s41592-024-02517-x
A proteome-wide quantitative platform for nanoscale spatially resolved extraction of membrane proteins into native nanodiscs
Abstract
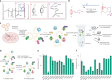
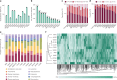
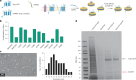
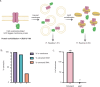
The native membrane environment profoundly influences every aspect of membrane protein (MP) biology. Despite this, the most prevalent method of studying MPs uses detergents to disrupt and remove this vital membrane context, impeding our ability to decipher the local molecular context and its effect. Here we develop a membrane proteome-wide platform that enables rapid spatially resolved extraction of target MPs directly from cellular membranes into native nanodiscs that maintain the local membrane context, using a library of membrane-active polymers. We accompany this with an open-access database that quantifies the polymer-specific extraction efficiency for 2,065 unique mammalian MPs and provides the most optimized extraction condition for each. To validate, we demonstrate how this resource can enable rapid extraction and purification of target MPs from different organellar membranes with high efficiency and purity. Further, we show how the database can be extended to capture overexpressed multiprotein complexes by taking two synaptic vesicle MPs. We expect these publicly available resources to empower researchers across disciplines to efficiently capture membrane 'nano-scoops' containing a target MP and interface with structural, functional and bioanalytical approaches.
© 2024. The Author(s).
Conflict of interest statement
Competing interests: The authors declare no competing interests.
Figures






Update of
-
A proteome-wide quantitative platform for nanoscale spatially resolved extraction of membrane proteins into native nanodiscs.bioRxiv [Preprint]. 2024 Aug 4:2024.02.10.579775. doi: 10.1101/2024.02.10.579775. bioRxiv. 2024. Update in: Nat Methods. 2025 Feb;22(2):412-421. doi: 10.1038/s41592-024-02517-x. PMID: 38405833 Free PMC article. Updated. Preprint.
References
MeSH terms
Substances
Grants and funding
- R35GM147095/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)
- R35 GM147095/GM/NIGMS NIH HHS/United States
- RM1GM149406/U.S. Department of Health & Human Services | NIH | National Institute of General Medical Sciences (NIGMS)
- R01 GM141192/GM/NIGMS NIH HHS/United States
- RM1 GM149406/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources

