Maintenance of niche architecture requires actomyosin and enables proper stem cell signaling and oriented division in the Drosophila testis
- PMID: 39620974
- PMCID: PMC11795290
- DOI: 10.1242/dev.204498
Maintenance of niche architecture requires actomyosin and enables proper stem cell signaling and oriented division in the Drosophila testis
Abstract
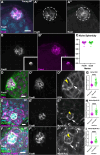
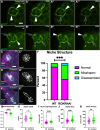
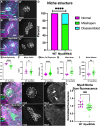
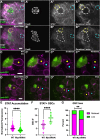
Stem cells are essential to repair and regenerate tissues, and often reside in a niche that controls their behavior. Here, we use the Drosophila testis niche, a paradigm for niche-stem cell interactions, to address the cell biological features that maintain niche structure and function during its steady-state operation. We report enrichment of Myosin II (MyoII) and a key regulator of actomyosin contractility (AMC), Rho Kinase (ROK), within the niche cell cortex at the interface with germline stem cells (GSCs). Compromising MyoII and ROK disrupts niche architecture, suggesting that AMC in niche cells is important to maintain its reproducible structure. Furthermore, defects in niche architecture disrupt GSC function. Our data suggest that the niche signals less robustly to adjacent germ cells yet permits increased numbers of cells to respond to the signal. Finally, compromising MyoII in niche cells leads to increased misorientation of centrosomes in GSCs as well as defects in the centrosome orientation checkpoint. Ultimately, this work identifies a crucial role for AMC-dependent maintenance of niche structure to ensure a proper complement of stem cells that correctly execute divisions.
Keywords: Drosophila; Cell division; Signaling; Stem cells.
© 2025. Published by The Company of Biologists.
Conflict of interest statement
Competing interests The authors declare no competing or financial interests.
Figures


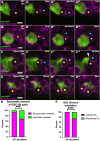

References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

