Opportunities and challenges of single-cell and spatially resolved genomics methods for neuroscience discovery
- PMID: 39627587
- PMCID: PMC11999325
- DOI: 10.1038/s41593-024-01806-0
Opportunities and challenges of single-cell and spatially resolved genomics methods for neuroscience discovery
Erratum in
-
Author Correction: Opportunities and challenges of single-cell and spatially resolved genomics methods for neuroscience discovery.Nat Neurosci. 2025 Jan;28(1):216. doi: 10.1038/s41593-024-01858-2. Nat Neurosci. 2025. PMID: 39681663 Free PMC article. No abstract available.
Abstract
Over the past decade, single-cell genomics technologies have allowed scalable profiling of cell-type-specific features, which has substantially increased our ability to study cellular diversity and transcriptional programs in heterogeneous tissues. Yet our understanding of mechanisms of gene regulation or the rules that govern interactions between cell types is still limited. The advent of new computational pipelines and technologies, such as single-cell epigenomics and spatially resolved transcriptomics, has created opportunities to explore two new axes of biological variation: cell-intrinsic regulation of cell states and expression programs and interactions between cells. Here, we summarize the most promising and robust technologies in these areas, discuss their strengths and limitations and discuss key computational approaches for analysis of these complex datasets. We highlight how data sharing and integration, documentation, visualization and benchmarking of results contribute to transparency, reproducibility, collaboration and democratization in neuroscience, and discuss needs and opportunities for future technology development and analysis.
© 2024. This is a U.S. Government work and not under copyright protection in the US; foreign copyright protection may apply.
Conflict of interest statement
Competing interests: S.A.L. declares a financial interest in AstronauTx and Synapticure. All other authors declare no competing interests.
Figures

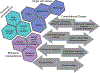


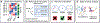
References
-
- Siletti K et al. Transcriptomic diversity of cell types across the adult human brain. Science 382, eadd7046 (2023). - PubMed
Publication types
MeSH terms
Grants and funding
- R01 NS123263/NS/NINDS NIH HHS/United States
- U01 AG072573/AG/NIA NIH HHS/United States
- R01 DA055823/DA/NIDA NIH HHS/United States
- R01 MH128364/MH/NIMH NIH HHS/United States
- ZIA NS003153/ImNIH/Intramural NIH HHS/United States
- MH103517/U.S. Department of Health & Human Services | NIH | National Institute of Mental Health (NIMH)
- P01 AG073082/AG/NIA NIH HHS/United States
- NS115821/U.S. Department of Health & Human Services | NIH | National Institute of Neurological Disorders and Stroke (NINDS)
- U01 DA052713/DA/NIDA NIH HHS/United States
- R01 MH126481/MH/NIMH NIH HHS/United States
- WT_/Wellcome Trust/United Kingdom
- R01 MH125516/MH/NIMH NIH HHS/United States
- MH126481/U.S. Department of Health & Human Services | NIH | National Institute of Mental Health (NIMH)
- UM1 HG012076/HG/NHGRI NIH HHS/United States
- R01 EY033353/EY/NEI NIH HHS/United States
- R01 NS126143/NS/NINDS NIH HHS/United States
- RF1 NS128908/NS/NINDS NIH HHS/United States
- RF1 AG079557/AG/NIA NIH HHS/United States
- R01MH128364/U.S. Department of Health & Human Services | NIH | National Institute of Mental Health (NIMH)
- R01 MH103517/MH/NIMH NIH HHS/United States
- R01MH125516/U.S. Department of Health & Human Services | NIH | National Institute of Mental Health (NIMH)
- RF1 NS126143/NS/NINDS NIH HHS/United States
- R01 AG079291/AG/NIA NIH HHS/United States
- HG011641/U.S. Department of Health & Human Services | NIH | National Human Genome Research Institute (NHGRI)
- NS126143/U.S. Department of Health & Human Services | NIH | National Institute of Neurological Disorders and Stroke (NINDS)
- U01 MH130962/MH/NIMH NIH HHS/United States
- UF1 NS115821/NS/NINDS NIH HHS/United States
- R01 HG011641/HG/NHGRI NIH HHS/United States
- R35 GM142889/GM/NIGMS NIH HHS/United States
LinkOut - more resources
Full Text Sources
Miscellaneous

