This is a preprint.
Revisiting the Plasmodium falciparum druggable genome using predicted structures and data mining
- PMID: 39649165
- PMCID: PMC11623766
- DOI: 10.21203/rs.3.rs-5412515/v1
Revisiting the Plasmodium falciparum druggable genome using predicted structures and data mining
Update in
-
Revisiting the Plasmodium falciparum druggable genome using predicted structures and data mining.NPJ Drug Discov. 2025;2(1):3. doi: 10.1038/s44386-025-00006-5. Epub 2025 Mar 4. NPJ Drug Discov. 2025. PMID: 40066064 Free PMC article.
Abstract
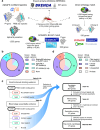
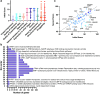
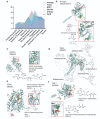
The identification of novel drug targets for the purpose of designing small molecule inhibitors is key component to modern drug discovery. In malaria parasites, discoveries of antimalarial targets have primarily occurred retroactively by investigating the mode of action of compounds found through phenotypic screens. Although this method has yielded many promising candidates, it is time- and resource-consuming and misses targets not captured by existing antimalarial compound libraries and phenotypic assay conditions. Leveraging recent advances in protein structure prediction and data mining, we systematically assessed the Plasmodium falciparum genome for proteins amenable to target-based drug discovery, identifying 867 candidate targets with evidence of small molecule binding and blood stage essentiality. Of these, 540 proteins showed strong essentiality evidence and lack inhibitors that have progressed to clinical trials. Expert review and rubric-based scoring of this subset based on additional criteria such as selectivity, structural information, and assay developability yielded 67 high priority candidates. This study also provides a genome-wide data resource and implements a generalizable framework for systematically evaluating and prioritizing novel pathogenic disease targets.
Keywords: Plasmodium blood stage targets; druggable genome; malaria data compendium.
Conflict of interest statement
Declarations Competing Interest Statement MKG has an equity interest in and is a cofounder and scientific advisor of VeraChem LLC, and is on the SABs of InCerebro Inc, Denovicon Therapeutics, and Beren Therapeutics. ELF and SPSR are employees of Novartis Pharma AG and may own shares in Novartis Pharma AG. KD holds stock in TropIQ Health Sciences. The rest of authors declare no competing interests.
Figures

References
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources

