Emerging mutation in SARS-CoV-2 facilitates escape from NK cell recognition and associates with enhanced viral fitness
- PMID: 39652590
- PMCID: PMC11658698
- DOI: 10.1371/journal.ppat.1012755
Emerging mutation in SARS-CoV-2 facilitates escape from NK cell recognition and associates with enhanced viral fitness
Abstract
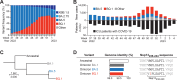
In addition to adaptive immunity, natural killer (NK) cells of the innate immune system contribute to the control of viral infections. The HLA-E-restricted SARS-CoV-2 Nsp13232-240 epitope VMPLSAPTL renders infected cells susceptible to NK cells by preventing binding to the inhibitory receptor NKG2A. Here, we report that a recently emerged methionine to isoleucine substitution at position 2 (pM2I) of Nsp13232-240 impairs binding of the mutated epitope to HLA-E and diminishes HLA-E/peptide complex stability. Structural analyses revealed altered occupancy of the HLA-E B-pocket as the underlying cause for reduced presentation and stability of the mutated epitope. Functionally, the reduced presentation of the mutated epitope correlated with elevated binding to NKG2A as well as with increased NK cell inhibition. Moreover, the pM2I mutation associated with enhanced estimated viral fitness and was transmitted to descendants of the SARS-CoV-2 BQ.1 variant. Interestingly, the mutated epitope resembles sequences of related peptides found in endemic common cold-causing human coronaviruses. Altogether, these findings indicate compromised peptide presentation as a viral adaptation to evade NK cell-mediated immunosurveillance by enabling enhanced presentation of self-peptide and restoring NKG2A-dependent inhibition of NK cells.
Copyright: © 2024 Bilev et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
H.-G.L. and Q.H. are consultants for and shareholders of Vycellix Inc., which had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Figures
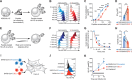



References
-
- Wilk AJ, Lee MJ, Wei B, Parks B, Pi R, Martinez-Colon GJ, et al.. Multi-omic profiling reveals widespread dysregulation of innate immunity and hematopoiesis in COVID-19. The Journal of experimental medicine. 2021;218(8). Epub 20210615. doi: 10.1084/jem.20210582 ; PubMed Central PMCID: PMC8210586. - DOI - PMC - PubMed
MeSH terms
Substances
Supplementary concepts
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

