Multiomics reveals tumor microenvironment remodeling in locally advanced gastric and gastroesophageal junction cancer following neoadjuvant immunotherapy and chemotherapy
- PMID: 39653554
- PMCID: PMC11629098
- DOI: 10.1136/jitc-2024-010041
Multiomics reveals tumor microenvironment remodeling in locally advanced gastric and gastroesophageal junction cancer following neoadjuvant immunotherapy and chemotherapy
Abstract
Background: Perioperative chemotherapy is the standard of care for patients with locally advanced gastric and gastroesophageal junction cancer. Recent evidence demonstrated the addition of programmed cell death protein 1 (PD-1) inhibitors enhanced therapeutic efficacy. However, the mechanisms of response and resistance remain largely undefined. A detailed multiomic investigation is essential to elucidate these mechanisms.
Methods: We performed whole-exome sequencing, whole-transcriptome sequencing, multiplex immunofluorescence and single-cell RNA sequencing on matched pretreatment and post-treatment samples from 30 patients enrolled in an investigator-initiated Phase 2 clinical trial (NCT04908566). All patients received neoadjuvant PD-1 inhibitors in combination with chemotherapy. A major pathologic response (MPR) was defined as the presence of no more than 10% residual viable tumor cells following treatment.
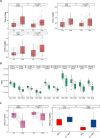
Results: Before treatment, the positive ratio of CD3+T cells in both the tumor parenchyma and stroma was significantly higher in the non-MPR group compared with the MPR group (p=0.042 and p=0.013, respectively). Least absolute shrinkage and selection operator regression was employed for feature gene selection and 13 genes were ultimately used to construct a predictive model for identifying MPR after surgery. The model exhibited a perfect area under curve (AUC) of 1.000 (95% CI: 1.000 to 1.000, p<0.001). Post-treatment analysis revealed a significant increase in CD3+T cells, CD8+T cells and NK cells in the tumor stroma of MPR patients. In the tumor parenchyma, aside from a marked increase in CD8+T cells and NK cells, a notable reduction in macrophage was also observed (all p<0.05). Importantly, forkheadbox protein 3 (FOXP3), the principal marker for regulatory T cells (Treg) cells, showed a significant decrease during treatment in MPR patients. FOXP3 expression in the non-MPR group was significantly higher than in the MPR group (p=0.0056) after treatment. Furthermore, single-cell RNA sequencing analysis confirmed that nearly all Treg cells were derived from the non-MPR group.
Conclusions: Our study highlights the critical role of dynamic changes within the tumor immune microenvironment in predicting the efficacy of neoadjuvant combined immunochemotherapy. We examined the disparities between MPR/non-MPR groups, shedding light on potential mechanisms of immune response and suppression. In addition to bolstering cytotoxic immune responses, specifically targeting Treg cells may be crucial for enhancing treatment outcomes.
Keywords: Gastric Cancer; Immunotherapy; Neoadjuvant; Tumor microenvironment - TME.
© Author(s) (or their employer(s)) 2024. Re-use permitted under CC BY-NC. No commercial re-use. See rights and permissions. Published by BMJ.
Conflict of interest statement
Competing interests: None declared.
Figures




References
-
- Al-Batran S-E, Homann N, Pauligk C, et al. Perioperative chemotherapy with fluorouracil plus leucovorin, oxaliplatin, and docetaxel versus fluorouracil or capecitabine plus cisplatin and epirubicin for locally advanced, resectable gastric or gastro-oesophageal junction adenocarcinoma (FLOT4): a randomised, phase 2/3 trial. Lancet. 2019;393:1948–57. doi: 10.1016/S0140-6736(18)32557-1. - DOI - PubMed
-
- Zhang X, Liang H, Li Z, et al. Perioperative or postoperative adjuvant oxaliplatin with S-1 versus adjuvant oxaliplatin with capecitabine in patients with locally advanced gastric or gastro-oesophageal junction adenocarcinoma undergoing D2 gastrectomy (RESOLVE): an open-label, superiority and non-inferiority, phase 3 randomised controlled trial. Lancet Oncol. 2021;22:1081–92. doi: 10.1016/S1470-2045(21)00297-7. - DOI - PubMed
Publication types
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
