Body mass index and fat influences the role of Bifidobacterium genus in lupus patients concerning fibrinogen levels
- PMID: 39654674
- PMCID: PMC11625790
- DOI: 10.3389/fmicb.2024.1471177
Body mass index and fat influences the role of Bifidobacterium genus in lupus patients concerning fibrinogen levels
Abstract
Introduction: Metabolic disorders and autoimmune diseases elicit distinct yet interconnected manifestations of inflammation, which may be boosted by an excess of body adiposity. The purpose of this investigation was to analyze anthropometric, biochemical, and inflammatory/coagulation variables concerning patients diagnosed with systemic lupus erythematosus (SLE) exploiting low-grade metabolic inflammation (MI), as reference.
Methods: A population stratification by body mass index (BMI), allowed to assess the impact of adiposity on the putative role of gut microbiota composition on coagulation markers. A total of 127 participants with MI and SLE were categorized into two main groups based on their BMI, following WHO criteria: a low BMI group (<30 kg/m2) and a high BMI group (≥30 kg/m2). Each group included recorded data on demographics, comorbidities, and key clinical markers. Anthropometric and body composition variables, clinical features, and inflammatory/coagulation markers were measured while fecal 16S rRNA sequencing was examined at the genus Bifidobacterium. Regression models were fitted to evaluate the relationship between gut microbiota, inflammatory/coagulation markers, and body weight in these types of diseases.
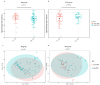
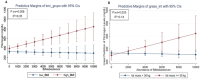
Results: The study revealed worse clinical outcomes in anthropometric, body composition, and clinical markers in low-grade MI conditions as compared to SLE. However, inflammatory and coagulation markers such as C-reactive protein (CRP) and fibrinogen were significantly more elevated in patients with SLE, which was exacerbated by high BMI/ body fat as compared to the other screened groups. An interaction analysis revealed that fibrinogen levels showed different trends when Bifidobacterium was increased depending on BMI/adiposity, which evidenced an effect modification by this microorganism in patients with SLE.
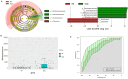
Discussion: These findings underline that gut microbiota composition, particularly the presence of Bifidobacterium, may play a crucial role in modulating inflammation and coagulation processes in patients with SLE and high fat. These insights highlight the potential of targeting gut microbiota as a therapeutic strategy to mitigate inflammation and improve clinical outcomes in SLE patients.
Keywords: Bifidobacterium; body mass index; fibrinogen; low-grade metabolic inflammation; systemic lupus erythematosus.
Copyright © 2024 Chero-Sandoval, Higuera-Gómez, Cuevas-Sierra, de Cuevillas, Castejón, Martínez-Urbistondo, Mellor-Pita, Moreno-Torres, de Luis and Martínez.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures
References
-
- Aringer M., Costenbader K., Daikh D., Brinks R., Mosca M., Ramsey-Goldman R., et al. (2019). 2019 European league against rheumatism/American College of Rheumatology Classification Criteria for systemic lupus erythematosus. Arthritis Rheumatol. 71, 1400–1412. doi: 10.1002/art.40930, PMID: - DOI - PMC - PubMed
-
- Ball D., Athanasiadou S. (2021). Understanding the interplay between the gut microbiome, nutrition and health. Proc. Nutr. Soc. 80, 383–385. doi: 10.1017/S0029665121003682 - DOI
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

