CD312 Promotes Paediatric Acute Lymphoblastic Leukaemia Through GNA15-Mediated Non-Classical GPCR Signalling Pathway
- PMID: 39656442
- PMCID: PMC11629795
- DOI: 10.1111/jcmm.70283
CD312 Promotes Paediatric Acute Lymphoblastic Leukaemia Through GNA15-Mediated Non-Classical GPCR Signalling Pathway
Erratum in
-
Correction to "CD312 Promotes Pediatric Acute Lymphoblastic Leukemia Through GNA15 Mediated Non-Classical GPCR Signaling Pathway".J Cell Mol Med. 2025 Jul;29(14):e70684. doi: 10.1111/jcmm.70684. J Cell Mol Med. 2025. PMID: 40702999 Free PMC article. No abstract available.
Abstract
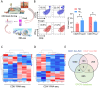
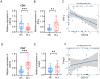
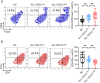
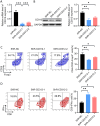
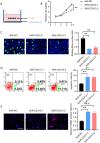
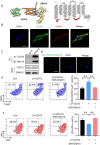
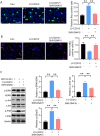
The bone marrow-infiltrated immune microenvironment plays a crucial role in blood system diseases, such as leukaemia. In this study, we aimed to investigate the critical role of the immune microenvironment in the onset and progression of childhood acute lymphoblastic leukaemia (ALL). Through high-throughput detection and screening of the GPCR database in the childhood ALL immune microenvironment, we identified CD312 as a candidate target. CD312 is associated with the distribution of Treg and CTL cells within the bone marrow immune microenvironment of ALL children. After CD312 knockdown, the proportion of the Treg subgroup in immune cells was significantly reduced, whereas the proportion of CTL subgroup cells was increased. CD312 exhibited good affinity with GNA15 in the transmembrane intracellular segment, and it could interact with GNA15. The BrdU staining assay revealed that the proliferation of leukaemia cells was enhanced in the CD312-overexpressed CD3+ T cells group via the phosphorylation of ERK, JNK and p38, whereas it was decreased by GNA15 knockdown in the co-culture system. In conclusion, our study suggests that CD312 fosters a suppressive immune microenvironment in the onset and progression of paediatric ALL through a GNA15-mediated non-classical GPCR signalling pathway.
Keywords: CD312; GPCR; acute lymphoblastic leukaemia; immune microenvironment.
© 2024 The Author(s). Journal of Cellular and Molecular Medicine published by Foundation for Cellular and Molecular Medicine and John Wiley & Sons Ltd.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
Unraveling the role of GPCR signaling in metabolic reprogramming and immune microenvironment of lung adenocarcinoma: a multi-omics study with experimental validation.Front Immunol. 2025 Jun 6;16:1606125. doi: 10.3389/fimmu.2025.1606125. eCollection 2025. Front Immunol. 2025. PMID: 40547013 Free PMC article.
-
Nutritional interventions for survivors of childhood cancer.Cochrane Database Syst Rev. 2016 Aug 22;2016(8):CD009678. doi: 10.1002/14651858.CD009678.pub2. Cochrane Database Syst Rev. 2016. PMID: 27545902 Free PMC article.
-
Hypothalamic-pituitary-adrenal (HPA) axis suppression after treatment with glucocorticoid therapy for childhood acute lymphoblastic leukaemia.Cochrane Database Syst Rev. 2017 Nov 6;11(11):CD008727. doi: 10.1002/14651858.CD008727.pub4. Cochrane Database Syst Rev. 2017. PMID: 29106702 Free PMC article.
-
Physical exercise training interventions for children and young adults during and after treatment for childhood cancer.Cochrane Database Syst Rev. 2016 Mar 31;3(3):CD008796. doi: 10.1002/14651858.CD008796.pub3. Cochrane Database Syst Rev. 2016. PMID: 27030386 Free PMC article.
-
Clinical effectiveness and cost-effectiveness of stem cell transplantation in the management of acute leukaemia: a systematic review.Health Technol Assess. 2010 Dec;14(54):iii-iv, ix-xi, 1-141. doi: 10.3310/hta14540. Health Technol Assess. 2010. PMID: 21138675
Cited by
-
GNA15 predicts poor outcomes as a novel biomarker related to M2 macrophage infiltration in ovarian cancer.Front Immunol. 2025 Feb 7;16:1512086. doi: 10.3389/fimmu.2025.1512086. eCollection 2025. Front Immunol. 2025. PMID: 39991148 Free PMC article.
References
-
- Hunger S. P., Tran T. H., Saha V., et al., “Dasatinib With Intensive Chemotherapy in de Novo Paediatric Philadelphia Chromosome‐Positive Acute Lymphoblastic Leukaemia (CA180‐372/COG AALL1122): A Single‐Arm, Multicentre, Phase 2 Trial,” Lancet Haematology 10 (2023): e510–e520. - PubMed
-
- Si Lim S. J., Ford J. B., and Hermiston M. L., “How I Treat Newly Diagnosed and Refractory T‐Cell Acute Lymphoblastic Lymphoma in Children and Young Adults,” Blood 141 (2023): 3019–3030. - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

