This is a preprint.
The SWIB domain-containing DNA topoisomerase I of Chlamydia trachomatis mediates DNA relaxation
- PMID: 39677648
- PMCID: PMC11642884
- DOI: 10.1101/2024.12.03.626651
The SWIB domain-containing DNA topoisomerase I of Chlamydia trachomatis mediates DNA relaxation
Update in
-
The SWIB domain-containing DNA topoisomerase I of Chlamydia trachomatis mediates DNA relaxation.J Bacteriol. 2025 Sep 18;207(9):e0019025. doi: 10.1128/jb.00190-25. Epub 2025 Aug 12. J Bacteriol. 2025. PMID: 40793755 Free PMC article.
Abstract
The obligate intracellular bacterial pathogen, Chlamydia trachomatis (Ct), has a distinct DNA topoisomerase I (TopA) with a C-terminal domain (CTD) homologous to eukaryotic SWIB domains. Despite the lack of sequence similarity at the CTDs between C. trachomatis TopA (CtTopA) and Escherichia coli TopA (EcTopA), full-length CtTopA removed negative DNA supercoils in vitro and complemented the growth defect of an E. coli topA mutant. We demonstrated that CtTopA is less processive in DNA relaxation than EcTopA in dose-response and time course studies. An antibody generated against the SWIB domain of CtTopA specifically recognized CtTopA but not EcTopA or Mycobacterium tuberculosis TopA (MtTopA), consistent with the sequence differences in their CTDs. The endogenous CtTopA protein is expressed at a relatively high level during the middle and late developmental stages of C. trachomatis. Conditional knockdown of topA expression using CRISPRi in C. trachomatis resulted in not only a developmental defect but also in the downregulation of genes linked to nucleotide acquisition from the host cells. Because SWIB-containing proteins are not found in prokaryotes beyond Chlamydia spp., these results imply a significant function for the SWIB-containing CtTopA in facilitating the energy metabolism of C. trachomatis for its unique intracellular growth.
Keywords: Chlamydia trachomatis; DNA relaxation; DNA topoisomerase I (TopA); SWIB domain; chlamydial developmental cycle; nucleotide metabolism; transcriptional regulation.
Figures



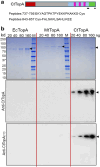


References
-
- Wang JC. 2002. Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol 3:430–40. - PubMed
Publication types
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous
