Identification of Ferroptosis-Related Gene in Age-Related Macular Degeneration Using Machine Learning
- PMID: 39679976
- PMCID: PMC11647999
- DOI: 10.1002/iid3.70059
Identification of Ferroptosis-Related Gene in Age-Related Macular Degeneration Using Machine Learning
Abstract
Background: Age-related macular degeneration (AMD) is a major cause of irreversible visual impairment, with dry AMD being the most prevalent form. Programmed cell death of retinal pigment epithelium (RPE) cells is a central mechanism in the pathogenesis of dry AMD. Ferroptosis, a recently identified form of programmed cell death, is characterized by iron accumulation-induced lipid peroxidation. This study aimed to investigate the involvement of ferroptosis in the progression of AMD.
Methods: A total of 41 samples of AMD and 50 normal samples were obtained from the data set GSE29801 for differential gene expression analysis and functional enrichment. Differentially expressed genes (DEGs) were selected and intersected with genes from the ferroptosis database to obtain differentially expressed ferroptosis-associated genes (DEFGs). Machine learning algorithms were employed to screen diagnostic genes. The diagnostic genes were subjected to Gene Set Enrichment Analysis (GSEA). Expression differences of diagnostic genes were validated in in vivo and in vitro models.
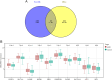
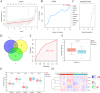
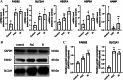
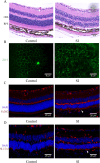
Results: We identified 462 DEGs when comparing normal and AMD samples. The GO enrichment analysis indicated significant involvement in key biological processes like collagen-containing extracellular matrix composition, positive cell adhesion regulation, and extracellular matrix organization. Through the intersection with ferroptosis gene sets, we pinpointed 10 DEFGs. Leveraging machine learning algorithms, we pinpointed five ferroptosis feature diagnostic genes: VEGFA, SLC2A1, HAMP, HSPB1, and FADS2. The subsequent experiments validated the increased expression of SLC2A1 and FADS2 in the AMD ferroptosis model.
Conclusion: The occurrence of ferroptosis could potentially contribute to the advancement of AMD. SLC2A1 and FADS2 have demonstrated promise as emerging diagnostic biomarkers and plausible therapeutic targets for AMD.
Keywords: age‐related macular degeneration; ferroptosis; machine learning.
© 2024 The Author(s). Immunity, Inflammation and Disease published by John Wiley & Sons Ltd.
Conflict of interest statement
The author declares no conflicts of interest.
Figures


References
-
- Wong W. L., Su X., Li X., et al., “Global Prevalence of Age‐Related Macular Degeneration and Disease Burden Projection for 2020 and 2040: a Systematic Review and Meta‐Analysis,” The Lancet Global Health 2, no. 2 (2014): e106–e116. - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

