Transcriptome immune-regulatory differences between leprosy patients and type 1 reaction patients, before onset of symptoms
- PMID: 39680574
- PMCID: PMC11684701
- DOI: 10.1371/journal.pntd.0011866
Transcriptome immune-regulatory differences between leprosy patients and type 1 reaction patients, before onset of symptoms
Abstract
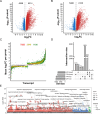
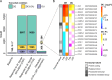
Leprosy is a chronic disease of the skin and peripheral nerves caused by Mycobacterium leprae. A major public health and clinical problem are leprosy reactions, which are inflammatory episodes that often contribute to nerve damage and disability. Type I reversal reactions (T1R) can occur after microbiological cure of leprosy and affect up to 50% of leprosy patients. Early intervention to prevent T1R and, hence, nerve damage, is a major focus of current leprosy control efforts. In a prospective study, we enrolled and collected samples from 32 leprosy patients before the onset of T1R. Whole blood aliquots were challenged with M. leprae sonicate or media and total RNA was extracted. After a three-year follow-up, the transcriptomic response was compared between cells from 22 patients who remained T1R-free and 10 patients who developed T1R during that period. Our analysis focused on differential transcript (i.e. isoform) expression and usage. Results showed that, at baseline, cells from T1R-destined and T1R-free subjects had no main difference in their transcripts expression and usage. However, the cells of T1R patients displayed a transcriptomic immune response to M. leprae antigens that was significantly different from the one of cells from leprosy patients who remained T1R-free. Transcripts with significantly higher upregulation in the T1R-destined group, compared to the cells from T1R-free patients, were enriched for pathways and GO terms involved in response to intracellular pathogens, apoptosis regulation and inflammatory processes. Similarly, transcript usage analysis pinpointed different transcript proportions in response to the in-vitro challenge of cells from T1R-destined patients. Hence, transcript usage in concert with transcript expression suggested a dysregulated inflammatory response including increased apoptosis regulation in the peripheral blood cells of T1R-destined patients before the onset of T1R symptoms. Combined, these results provided detailed insight into the pathogenesis of T1R.
Copyright: © 2024 Correa-Macedo et al. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Conflict of interest statement
The authors have declared that no competing interests exist.
Figures



Similar articles
-
Gene set signature of reversal reaction type I in leprosy patients.PLoS Genet. 2013;9(7):e1003624. doi: 10.1371/journal.pgen.1003624. Epub 2013 Jul 11. PLoS Genet. 2013. PMID: 23874223 Free PMC article.
-
Longitudinal immune responses and gene expression profiles in type 1 leprosy reactions.J Clin Immunol. 2014 Feb;34(2):245-55. doi: 10.1007/s10875-013-9979-x. Epub 2013 Dec 27. J Clin Immunol. 2014. PMID: 24370984
-
Autophagy-Associated IL-15 Production Is Involved in the Pathogenesis of Leprosy Type 1 Reaction.Cells. 2021 Aug 27;10(9):2215. doi: 10.3390/cells10092215. Cells. 2021. PMID: 34571865 Free PMC article.
-
Type 1 reaction in leprosy: a model for a better understanding of tissue immunity under an immunopathological condition.Expert Rev Clin Immunol. 2015 Mar;11(3):391-407. doi: 10.1586/1744666X.2015.1012501. Epub 2015 Feb 10. Expert Rev Clin Immunol. 2015. PMID: 25666357 Review.
-
Correlates of immune exacerbations in leprosy.Semin Immunol. 2018 Oct;39:111-118. doi: 10.1016/j.smim.2018.06.003. Epub 2018 Jun 24. Semin Immunol. 2018. PMID: 29950273 Review.
References
-
- World Health Organization = Organisation mondiale de la S. Global leprosy (Hansen disease) update, 2020: impact of COVID-19 on global leprosy control–Situation de la lèpre (maladie de Hansen) dans le monde, 2020: impact de la COVID-19 sur les activités mondiales de lutte contre la lèpre. Weekly Epidemiological Record = Relevé épidémiologique hebdomadaire. 2021;96(36):421–44.
-
- Teles RMB, Lu J, Tió-Coma M, Goulart IMB, Banu S, Hagge D, et al.. Identification of a systemic interferon-γ inducible antimicrobial gene signature in leprosy patients undergoing reversal reaction. PLoS Negl Trop Dis. 2019;13(10):e0007764. Epub 2019/10/11. doi: 10.1371/journal.pntd.0007764 ; PubMed Central PMCID: PMC6805014. - DOI - PMC - PubMed
MeSH terms
LinkOut - more resources
Full Text Sources
Medical

