Genome Instability and Senescence Are Markers of Cornelia de Lange Syndrome Cells
- PMID: 39682772
- PMCID: PMC11640591
- DOI: 10.3390/cells13232025
Genome Instability and Senescence Are Markers of Cornelia de Lange Syndrome Cells
Abstract
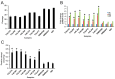
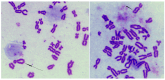
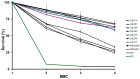
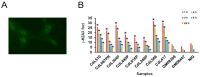
Cornelia de Lange syndrome (CdLS) is a rare, dominantly inherited multisystem developmental disorder. Pathogenic variants in genes encoding the structural subunits and regulatory proteins of the cohesin complex (NIPBL, SMC1A, SMC3, HDAC8, and RAD21) are the primary contributors to the pathogenesis of CdLS. Pathogenic variations in these genes disrupt normal cohesin function, leading to the syndrome's diverse and complex clinical presentation. In this study, we discovered that cells harboring variants in the NIPBL, SMC1A and HDAC8 genes exhibit spontaneous genome instability, elevated oxidative stress and premature cellular aging. These findings suggest that cohesin plays a critical role in maintaining proper cellular function and highlight its contribution to the pathophysiology seen in the related diagnoses.
Keywords: Cornelia de Lange syndrome; HDAC8; NIPBL; SMC1A; cohesin; genome instability; oxidative stress; senescence.
Conflict of interest statement
The authors declare no conflicts of interest. The funders had no role in the design of the study; in the collection, analyses, or interpretation of data; in the writing of the manuscript; or in the decision to publish the results.
Figures
References
Publication types
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

