Synthesis and Physiochemical Properties of Sulphated Tamarind (Tamarindus indica L.) Seed Polysaccharide
- PMID: 39683670
- PMCID: PMC11643769
- DOI: 10.3390/molecules29235510
Synthesis and Physiochemical Properties of Sulphated Tamarind (Tamarindus indica L.) Seed Polysaccharide
Abstract
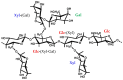
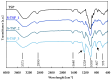
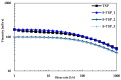
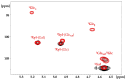
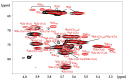
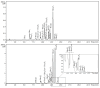
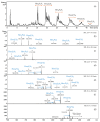
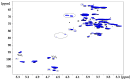
Tamarind seed polysaccharide (TSP) is a neutral water-soluble galactoxyloglucan isolated from the seed kernel of Tamarindus indica with average molecular weight (Mw) 600-800 kDa. The high viscosity of TSP slows solubilisation, and the absence of charged substituent hinders the formation of electrostatic interactions with biomolecules. TSP was sulphated in a one-step process using dimethylformamide as a solvent, and sulphur trioxide-pyridine complex as a sulphating reagent. Studies of chemical structure, molecular weight distribution and viscosity were conducted to characterise the synthesised products. The sulphation degree was established by conductimetric titration; the sulphate group distribution was studied by NMR spectroscopy and liquid chromatography-mass spectrometry, and sulphated TSP oligomers were obtained by enzymatic degradation with cellulase and/or xyloglucanase. Sulphated products showed higher solubility than TSP, Mws in the range of 700-1000 kDa, a sulphation degree of two to four per subunit and pseudoplastic behaviour. A preliminary study of mucoadhesion revealed the unexpected interaction of S-TSP with mucin, providing a route by which sulphated TSP interactions with biomolecules may be influenced.
Keywords: LC-MS; NMR; enzymatic hydrolysis; molecular weight; structural characterization; sulphated polysaccharide; tamarind seed polysaccharide; viscosity.
Conflict of interest statement
Authors Sabrina Ziliani, Fabio Neggiani and Marco Sansò were employed by the company Istituto Farmaco Biologico Sperimentale Srl. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Figures


Similar articles
-
Composition and Rheological Properties of Polysaccharide Extracted from Tamarind (Tamarindus indica L.) Seed.Molecules. 2019 Mar 28;24(7):1218. doi: 10.3390/molecules24071218. Molecules. 2019. PMID: 30925745 Free PMC article.
-
An amendment to the fine structure of galactoxyloglucan from Tamarind (Tamarindus indica L.) seed.Int J Biol Macromol. 2020 Apr 15;149:1189-1197. doi: 10.1016/j.ijbiomac.2020.01.284. Epub 2020 Jan 30. Int J Biol Macromol. 2020. PMID: 32006579
-
Screening of polysaccharides from tamarind, fenugreek and jackfruit seeds as pharmaceutical excipients.Int J Biol Macromol. 2015 Aug;79:756-60. doi: 10.1016/j.ijbiomac.2015.05.018. Epub 2015 May 22. Int J Biol Macromol. 2015. PMID: 26007663
-
State-of-the-art progress on tamarind seed polysaccharide (Tamarindus indica) and its diverse potential applications, a review with insight.Carbohydr Polym. 2024 May 1;331:121847. doi: 10.1016/j.carbpol.2024.121847. Epub 2024 Feb 1. Carbohydr Polym. 2024. PMID: 38388032 Review.
-
Tamarind (Tamarindus indica L.) Seed Polysaccharide: A promising biopolymer for drug delivery, wound healing, tissue engineering and beyond.Carbohydr Res. 2025 Jun;552:109454. doi: 10.1016/j.carres.2025.109454. Epub 2025 Mar 4. Carbohydr Res. 2025. PMID: 40056791 Review.
Cited by
-
Preparation, structural characterization and health benefits of polysaccharides from Rubus chingii Hu: a review.Front Pharmacol. 2025 May 9;16:1592046. doi: 10.3389/fphar.2025.1592046. eCollection 2025. Front Pharmacol. 2025. PMID: 40417219 Free PMC article. Review.
References
-
- Shukla A.K., Bishnoi R.S., Kumar M., Fenin V., Jain C.P. Applications of Tamarind seeds Polysaccharide-based copolymers in Controlled Drug Delivery: An overview. Asian J. Pharm. Pharmacol. 2018;4:23–30. doi: 10.31024/ajpp.2018.4.1.5. - DOI
-
- Ren L., Yang Y., Bian X., Li X., Wang B., Wang D., Su D., Liu L., Yu D., Guo X., et al. Physicochemical, Rheological, Structural, Antioxidant, and Antimicrobial Properties of Polysaccharides Extracted from Tamarind Seeds. J. Food Qual. 2022;2022:9788248. doi: 10.1155/2022/9788248. - DOI
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Miscellaneous

