Calcium Indicators with Fluorescence Lifetime-Based Signal Readout: A Structure-Function Study
- PMID: 39684209
- PMCID: PMC11640911
- DOI: 10.3390/ijms252312493
Calcium Indicators with Fluorescence Lifetime-Based Signal Readout: A Structure-Function Study
Abstract
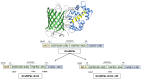
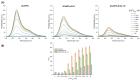
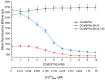
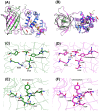
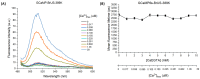
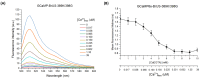
The calcium cation is a crucial signaling molecule involved in numerous cellular pathways. Beyond its role as a messenger or modulator in intracellular cascades, calcium's function in excitable cells, including nerve impulse transmission, is remarkable. The central role of calcium in nervous activity has driven the rapid development of fluorescent techniques for monitoring this cation in living cells. Specifically, genetically encoded calcium indicators (GECIs) are the most in-demand molecular tools in their class. In this work, we address two issues of calcium imaging by designing indicators based on the successful GCaMP6 backbone and the fluorescent protein BrUSLEE. The first indicator variant (GCaMP6s-BrUS), with a reduced, calcium-insensitive fluorescence lifetime, has potential in monitoring calcium dynamics with a high temporal resolution in combination with advanced microscopy techniques, such as light beads microscopy, where the fluorescence lifetime limits acquisition speed. Conversely, the second variant (GCaMP6s-BrUS-145), with a flexible, calcium-sensitive fluorescence lifetime, is relevant for static measurements, particularly for determining absolute calcium concentration values using fluorescence lifetime imaging microscopy (FLIM). To identify the structural determinants of calcium sensitivity in these indicator variants, we determine their spatial structures. A comparative structural analysis allowed the optimization of the GCaMP6s-BrUS construct, resulting in an indicator variant combining calcium-sensitive behavior in the time domain and enhanced molecular brightness. Our data may serve as a starting point for further engineering efforts towards improved GECI variants with fine-tuned fluorescence lifetimes.
Keywords: FLIM; fluorescence lifetime; genetically encoded indicators; quantitative calcium imaging; structural analysis.
Conflict of interest statement
The authors declare no conflicts of interest.
Figures
Similar articles
-
A Bioluminescent Ca2+ Indicator Based on a Topological Variant of GCaMP6s.Chembiochem. 2019 Feb 15;20(4):516-520. doi: 10.1002/cbic.201800255. Epub 2018 Jul 31. Chembiochem. 2019. PMID: 29934970
-
cNTnC and fYTnC2, Genetically Encoded Green Calcium Indicators Based on Troponin C from Fast Animals.Int J Mol Sci. 2022 Nov 23;23(23):14614. doi: 10.3390/ijms232314614. Int J Mol Sci. 2022. PMID: 36498942 Free PMC article.
-
Crystal Structure of Bright Fluorescent Protein BrUSLEE with Subnanosecond Fluorescence Lifetime; Electric and Dynamic Properties.Int J Mol Sci. 2023 Mar 29;24(7):6403. doi: 10.3390/ijms24076403. Int J Mol Sci. 2023. PMID: 37047378 Free PMC article.
-
New red-fluorescent calcium indicators for optogenetics, photoactivation and multi-color imaging.Biochim Biophys Acta. 2014 Oct;1843(10):2284-306. doi: 10.1016/j.bbamcr.2014.03.010. Epub 2014 Mar 27. Biochim Biophys Acta. 2014. PMID: 24681159 Review.
-
[Microscopic imaging of calcium ions with genetically encoded calcium indicators].Postepy Biochem. 2017;63(1):34-43. Postepy Biochem. 2017. PMID: 28409573 Review. Polish.
Cited by
-
MicroRNA: unveiling novel mechanistic and theranostic pathways in diabetic cardiomyopathy.Front Pharmacol. 2025 Jul 23;16:1613844. doi: 10.3389/fphar.2025.1613844. eCollection 2025. Front Pharmacol. 2025. PMID: 40771931 Free PMC article. Review.
References
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources

