Identification and development of TRPM4 antagonists to counteract neuronal excitotoxicity
- PMID: 39687019
- PMCID: PMC11648915
- DOI: 10.1016/j.isci.2024.111425
Identification and development of TRPM4 antagonists to counteract neuronal excitotoxicity
Abstract
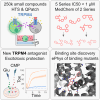
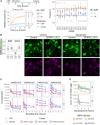
Neurodegeneration in central nervous system disorders is linked to dysregulated neuronal calcium. Direct inhibition of glutamate-induced neuronal calcium influx, particularly via N-methyl-D-aspartate receptors (NMDAR), has led to adverse effects and clinical trial failures. A more feasible approach is to modulate NMDAR activity or calcium signaling indirectly. In this respect, the calcium-activated non-selective cation channel transient receptor potential melastatin 4 (TRPM4) has been identified as a promising target. However, high affinity and specific antagonists are lacking. Here, we conducted high-throughput screening of a compound library to identify high affinity TRPM4 antagonists. This yielded five lead compound series with nanomolar half-maximal inhibitory concentration values. Through medicinal chemistry optimization of two series, we established detailed structure-activity relationships and inhibition of excitotoxicity in neurons. Moreover, we identified their potential binding site supported by electrophysiological measurements. These potent TRPM4 antagonists are promising drugs for treating neurodegenerative disorders and TRPM4-related pathologies, potentially overcoming previous therapeutic challenges.
Keywords: Molecular biology; Molecular neuroscience; Neuroscience.
© 2024 The Author(s).
Conflict of interest statement
L.B-.L. and M.A.F. are inventors on a filed patent application covering the therapeutic use of the described compounds to block TRPM4. All other authors declare no conflicts of interest.
Figures



References
LinkOut - more resources
Full Text Sources

