Increased CSN5 expression enhances the sensitivity to lenalidomide in multiple myeloma cells
- PMID: 39687025
- PMCID: PMC11647120
- DOI: 10.1016/j.isci.2024.111399
Increased CSN5 expression enhances the sensitivity to lenalidomide in multiple myeloma cells
Abstract
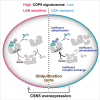
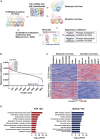
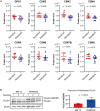
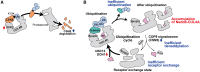
Lenalidomide (LEN) is commonly used as an effective therapeutic agent for multiple myeloma (MM). However, in some patients, primary resistance to LEN is observed, the mechanisms of which remain poorly understood. In this study, we combined a LEN sensitivity assay with proteomics data from 15 MM cell lines to identify protein expression profiles associated with primary LEN resistance. Our findings revealed that CSN5 expression is lower in LEN-resistant cell lines than in LEN-sensitive lines. Moreover, we established that CSN5 is degraded via the cullin-RING ubiquitin ligase (CRL)-mediated ubiquitin-proteasome pathway through ubiquitination at lysine 194. Our data suggest that reduced CSN5 expression leads to abnormalities in the ubiquitination cycle of CRL4A, resulting in the inhibition of LEN-mediated degradation of IKZF1 and IKZF3. These findings delineate an additional mechanism of LEN resistance in MM cells and may contribute to the development of alternative therapeutic strategies to overcome LEN resistance.
Keywords: Cancer; Cell biology; Molecular biology; Proteomics.
© 2024 The Authors.
Conflict of interest statement
The authors declare no competing interests.
Figures



Similar articles
-
SUMOylation inhibition enhances multiple myeloma sensitivity to lenalidomide.Cancer Gene Ther. 2023 Apr;30(4):567-574. doi: 10.1038/s41417-022-00450-9. Epub 2022 Mar 25. Cancer Gene Ther. 2023. PMID: 35338347 Free PMC article.
-
Lys-63-specific deubiquitinase BRCC36 enhances the sensitivity of multiple myeloma cells to lenalidomide by inhibiting lysosomal degradation of cereblon.Cell Mol Life Sci. 2024 Aug 13;81(1):349. doi: 10.1007/s00018-024-05390-1. Cell Mol Life Sci. 2024. PMID: 39136771 Free PMC article.
-
Caspase-8 Inhibition Prevents the Cleavage and Degradation of E3 Ligase Substrate Receptor Cereblon and Potentiates Its Biological Function.Front Cell Dev Biol. 2020 Dec 17;8:605989. doi: 10.3389/fcell.2020.605989. eCollection 2020. Front Cell Dev Biol. 2020. PMID: 33392195 Free PMC article.
-
[Molecular Mechanism of CRBN in the Activity of Lenalidomid eagainst Myeloma--Review].Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2018 Aug;26(4):1240-1243. doi: 10.7534/j.issn.1009-2137.2018.04.050. Zhongguo Shi Yan Xue Ye Xue Za Zhi. 2018. PMID: 30111438 Review. Chinese.
-
Lenalidomide in Multiple Myeloma: Review of Resistance Mechanisms, Current Treatment Strategies and Future Perspectives.Cancers (Basel). 2023 Feb 2;15(3):963. doi: 10.3390/cancers15030963. Cancers (Basel). 2023. PMID: 36765919 Free PMC article. Review.
References
-
- Rajkumar S.V., Dimopoulos M.A., Palumbo A., Blade J., Merlini G., Mateos M.V., Kumar S., Hillengass J., Kastritis E., Richardson P., et al. International Myeloma Working Group updated criteria for the diagnosis of multiple myeloma. Lancet Oncol. 2014;15:e538–e548. doi: 10.1016/S1470-2045(14)70442-5. - DOI - PubMed
LinkOut - more resources
Full Text Sources

