Quantifying conformational changes in the TCR:pMHC-I binding interface
- PMID: 39687625
- PMCID: PMC11646856
- DOI: 10.3389/fimmu.2024.1491656
Quantifying conformational changes in the TCR:pMHC-I binding interface
Abstract
Background: T cells form one of the key pillars of adaptive immunity. Using their surface bound T cell antigen receptors (TCRs), these cells screen millions of antigens presented by major histocompatibility complex (MHC) or MHC-like molecules. In other protein families, the dynamics of protein-protein interactions have important implications for protein function. Case studies of TCR:class I peptide-MHCs (pMHC-Is) structures have reported mixed results on whether the binding interfaces undergo conformational change during engagement and no robust statistical quantification has been done to generalise these results. Thus, it remains an open question of whether movement occurs in the binding interface that enables the recognition and activation of T cells.
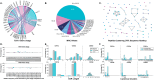
Methods: In this work, we quantify the conformational changes in the TCR:pMHC-I binding interface by creating a dataset of 391 structures, comprising 22 TCRs, 19 MHC alleles, and 79 peptide structures in both unbound (apo) and bound (holo) conformations.
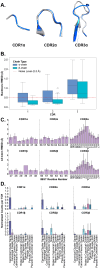
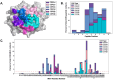
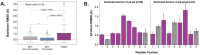
Results: In support of some case studies, we demonstrate that all complementarity determining region (CDR) loops move to a certain extent but only CDR3α and CDR3β loops modify their shape when binding pMHC-Is. We also map the contacts between TCRs and pMHC-Is, generating a novel fingerprint of TCRs on MHC molecules and show that the CDR3α tends to bind the N-terminus of the peptide and the CDR3β tends to bind the C-terminus of the peptide. Finally, we show that the presented peptides can undergo conformational changes when engaged by TCRs, as has been reported in past literature, but novelly show these changes depend on how the peptides are anchored in the MHC binding groove.
Conclusions: Our work has implications in understanding the behaviour of TCR:pMHC-I interactions and providing insights that can be used for modelling Tcell antigen specificity, an ongoing grand challenge in immunology.
Keywords: HLA; MHC; T cell antigen specificity; TCR; conformational changes; peptide; structural biology.
Copyright © 2024 McMaster, Thorpe, Rossjohn, Deane and Koohy.
Conflict of interest statement
The authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest. The author(s) declared that they were an editorial board member of Frontiers, at the time of submission. This had no impact on the peer review process and the final decision.
Figures

Similar articles
-
T-cell receptor (TCR)-peptide specificity overrides affinity-enhancing TCR-major histocompatibility complex interactions.J Biol Chem. 2014 Jan 10;289(2):628-38. doi: 10.1074/jbc.M113.522110. Epub 2013 Nov 6. J Biol Chem. 2014. PMID: 24196962 Free PMC article.
-
TCR scanning of peptide/MHC through complementary matching of receptor and ligand molecular flexibility.J Immunol. 2014 Mar 15;192(6):2885-91. doi: 10.4049/jimmunol.1302953. Epub 2014 Feb 12. J Immunol. 2014. PMID: 24523505 Free PMC article.
-
Molecular basis of a dominant T cell response to an HIV reverse transcriptase 8-mer epitope presented by the protective allele HLA-B*51:01.J Immunol. 2014 Apr 1;192(7):3428-34. doi: 10.4049/jimmunol.1302667. Epub 2014 Mar 5. J Immunol. 2014. PMID: 24600035 Free PMC article.
-
Conformational changes and flexibility in T-cell receptor recognition of peptide-MHC complexes.Biochem J. 2008 Oct 15;415(2):183-96. doi: 10.1042/BJ20080850. Biochem J. 2008. PMID: 18800968 Free PMC article. Review.
-
A structural voyage toward an understanding of the MHC-I-restricted immune response: lessons learned and much to be learned.Immunol Rev. 2012 Nov;250(1):61-81. doi: 10.1111/j.1600-065X.2012.01159.x. Immunol Rev. 2012. PMID: 23046123 Review.
Cited by
-
T-cell receptor structures and predictive models reveal comparable alpha and beta chain structural diversity despite differing genetic complexity.Commun Biol. 2025 Mar 4;8(1):362. doi: 10.1038/s42003-025-07708-6. Commun Biol. 2025. PMID: 40038394 Free PMC article.
References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

