NEDD4L mediates intestinal epithelial cell ferroptosis to restrict inflammatory bowel diseases and colorectal tumorigenesis
- PMID: 39688910
- PMCID: PMC11785928
- DOI: 10.1172/JCI173994
NEDD4L mediates intestinal epithelial cell ferroptosis to restrict inflammatory bowel diseases and colorectal tumorigenesis
Abstract
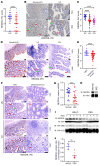
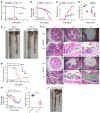
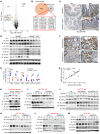
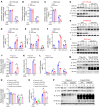
Various factors play key roles in maintaining intestine homeostasis. Disruption of the balance may lead to inflammatory bowel diseases and even colorectal cancer (CRC). Loss or gain of function of many key proteins can result in dysregulated intestinal homeostasis. Our research demonstrated that neural precursor cells expressed developmentally downregulated 4-like protein (NEDD4L, or NEDD4-2), a type of HECT family E3 ubiquitin ligase, played an important role in maintaining intestinal homeostasis. NEDD4L expression was significantly inhibited in intestinal epithelial cells (IECs) of patients with Crohn's disease, ulcerative colitis, and CRC. Global KO of NEDD4L or its deficiency in IECs exacerbated colitis induced by dextran sulfate sodium (DSS) and 2,4,6-trinitrobenzene sulfonic acid (TNBS) and CRC induced by azoxymethane and DSS. Mechanistically, NEDD4L deficiency in IECs inhibited expression of the key ferroptosis regulator glutathione peroxidase 4 (GPX4) by reducing the protein expression of solute carrier family 3 member 2 (SLC3A2) without affecting its gene expression, ultimately promoting DSS-induced IEC ferroptosis. Importantly, ferroptosis inhibitors reduced the susceptibility of NEDD4L-deficient mice to colitis and colitis-associated CRC. Thus, NEDD4L is an important regulator in IEC ferroptosis, maintaining intestinal homeostasis, making it a potential clinical target for diagnosing and treating IBDs.
Keywords: Cell biology; Epithelial transport of ions and water; Inflammation; Inflammatory bowel disease; Molecular biology.
Figures


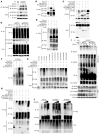



References
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Medical
Molecular Biology Databases
Research Materials

