DDO-adjuvanted influenza A virus nucleoprotein mRNA vaccine induces robust humoral and cellular type 1 immune responses and protects mice from challenge
- PMID: 39692514
- PMCID: PMC11796404
- DOI: 10.1128/mbio.03589-24
DDO-adjuvanted influenza A virus nucleoprotein mRNA vaccine induces robust humoral and cellular type 1 immune responses and protects mice from challenge
Abstract
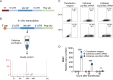
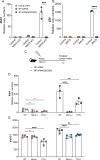
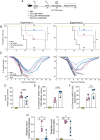
A challenge in viral vaccine development is to produce vaccines that generate both neutralizing antibodies to prevent infection and cytotoxic CD8+ T-cells that target conserved viral proteins and can eliminate infected cells to control virus spread. mRNA technology offers an opportunity to design vaccines based on conserved CD8-targeting epitopes, but achieving robust antigen-specific CD8+ T-cells remains a challenge. Here, we tested the viral-derived oligonucleotide DDO268 as an adjuvant in the context of a model influenza A virus (IAV) nucleoprotein (NP) mRNA vaccine in C57BL/6 mice. DDO268 when co-packaged with mRNA in lipid nanoparticles is sensed by RIG I-like receptors and safely induces local type I interferon (IFN) production followed by dendritic cells type 1 activation and migration to the draining lymph nodes. This early response triggered by DDO268 improved the generation of IgG2c antibodies and antigen-specific Th1 CD4+ and CD8+ T-cells (IFNγ+TNFα+IL2+) that provided enhanced protection against lethal IAV challenge. In addition, the inclusion of DDO268 reduced the antigen dose required to achieve protection. These results highlight the potential of DDO268 as an effective mRNA vaccine adjuvant and show that an IAV NP mRNA/DDO268 vaccine is a promising approach for generating protective immunity against conserved internal IAV epitopes.IMPORTANCEVaccines that generate neutralizing antibodies and cytotoxic CD8+ T-cells targeting conserved epitopes are ideal for effective protection against viruses. mRNA vaccines combined with the right adjuvant offer a promising solution to this challenge. We show that the virus-derived oligonucleotide DDO268 enhances antibody and T-cell responses to an influenza A virus (IAV) nucleoprotein mRNA vaccine in mice. DDO268 safely induces local type I interferon production and stimulates dendritic cell activation providing enhanced protection against IAV challenge. In addition, the adjuvant activity of DDO268 allows for the use of lower antigen doses during vaccination.
Keywords: adjuvants; mRNA vaccine; type I IFN.
Conflict of interest statement
C.B.L. and V.G. have provisional filed patent applications related to the manuscript. The University of Pennsylvania and C.B.L. have a patent for Methods and Compositions for Stimulating Immune Response Using Potent Immunostimulatory RNA Motifs.
Figures





Update of
-
DDO-adjuvanted influenza A virus nucleoprotein mRNA vaccine induces robust humoral and cellular type 1 immune responses and protects mice from challenge.bioRxiv [Preprint]. 2024 Oct 29:2024.10.27.620508. doi: 10.1101/2024.10.27.620508. bioRxiv. 2024. Update in: mBio. 2025 Feb 05;16(2):e0358924. doi: 10.1128/mbio.03589-24. PMID: 39553933 Free PMC article. Updated. Preprint.
References
-
- Freyn AW, Ramos da Silva J, Rosado VC, Bliss CM, Pine M, Mui BL, Tam YK, Madden TD, de Souza Ferreira LC, Weissman D, Krammer F, Coughlan L, Palese P, Pardi N, Nachbagauer R. 2020. A multi-targeting, nucleoside-modified mRNA influenza virus vaccine provides broad protection in mice. Mol Ther 28:1569–1584. doi: 10.1016/j.ymthe.2020.04.018 - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Medical
Research Materials
Miscellaneous

