Integrative multi-omics analysis reveals the contribution of neoVTX genes to venom diversity of Synanceia verrucosa
- PMID: 39695923
- PMCID: PMC11657881
- DOI: 10.1186/s12864-024-11149-6
Integrative multi-omics analysis reveals the contribution of neoVTX genes to venom diversity of Synanceia verrucosa
Abstract
Background: Animal venom systems are considered as valuable model for investigating the molecular mechanisms underlying phenotypic evolution. Stonefish are the most venomous and dangerous fish because of severe human envenomation and occasionally fatalities, whereas the genomic background of their venom has not been fully explored compared with that in other venomous animals.
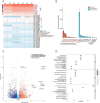
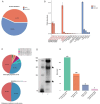
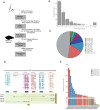
Results: In this study, we followed modern venomic pipelines to decode the Synanceia verrucosa venom components. A catalog of 478 toxin genes was annotated based on our assembled chromosome-level genome. Integrative analysis of the high-quality genome, the transcriptome of the venom gland, and the proteome of crude venom revealed mechanisms underlying the venom complexity in S. verrucosa. Six tandem-duplicated neoVTX subunit genes were identified as the major source for the neoVTX protein production. Further isoform sequencing revealed massive alternative splicing events with a total of 411 isoforms demonstrated by the six genes, which further contributed to the venom diversity. We then characterized 12 dominantly expressed toxin genes in the venom gland, and 11 of which were evidenced to produce the venom protein components, with the neoVTX proteins as the most abundant. Other major venom proteins included a presumed CRVP, Kuntiz-type serine protease inhibitor, calglandulin protein, and hyaluronidase. Besides, a few of highly abundant non-toxin proteins were also characterized and they were hypothesized to function in housekeeping or hemostasis maintaining roles in the venom gland. Notably, gastrotropin like non-toxin proteins were the second highest abundant proteins in the venom, which have not been reported in other venomous animals and contribute to the unique venom properties of S. verrucosa.
Conclusions: The results identified the major venom composition of S. verrucosa, and highlighted the contribution of neoVTX genes to the diversity of venom composition through tandem-duplication and alternative splicing. The diverse neoVTX proteins in the venom as lethal particles are important for understanding the adaptive evolution of S. verrucosa. Further functional studies are encouraged to exploit the venom components of S. verrucosa for pharmaceutical innovation.
Keywords: Synanceia verrucosa; Alternative splicing; Multi-omics; Tandem duplication; Venom diversity.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: All the experiments were conducted in accordance with guidelines approved by the Animal Ethical and Welfare Committee of Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou). Consent for publication: Not applicable. Competing interests: The authors declare no competing interests.
Figures

Similar articles
-
Purification, properties and cDNA cloning of neoverrucotoxin (neoVTX), a hemolytic lethal factor from the stonefish Synanceia verrucosa venom.Biochim Biophys Acta. 2006 Nov;1760(11):1713-22. doi: 10.1016/j.bbagen.2006.08.017. Epub 2006 Aug 26. Biochim Biophys Acta. 2006. PMID: 17023116
-
A Chromosome-Level Genome Assembly of the Reef Stonefish (Synanceia verrucosa) Provides Novel Insights into Stonustoxin (sntx) Genes.Mol Biol Evol. 2023 Oct 4;40(10):msad215. doi: 10.1093/molbev/msad215. Mol Biol Evol. 2023. PMID: 37770059 Free PMC article.
-
Properties and cDNA cloning of a hyaluronidase from the stonefish Synanceia verrucosa venom.Toxicon. 2011 Sep 15;58(4):285-92. doi: 10.1016/j.toxicon.2011.07.014. Epub 2011 Jul 28. Toxicon. 2011. PMID: 21820461
-
[Pharmacological properties of fish venoms].C R Seances Soc Biol Fil. 1998;192(3):503-48. C R Seances Soc Biol Fil. 1998. PMID: 9759386 Review. French.
-
The Geographic Distribution, Venom Components, Pathology and Treatments of Stonefish (Synanceia spp.) Venom.Mar Drugs. 2021 May 24;19(6):302. doi: 10.3390/md19060302. Mar Drugs. 2021. PMID: 34073964 Free PMC article. Review.
References
-
- Zancolli G, Casewell NR. Venom systems as models for studying the origin and regulation of evolutionary novelties. Mol Biol Evol. 2020;37(10):2777–90. - PubMed
-
- Suranse V, Srikanthan A, Sunagar K. Animal Venoms: Origin, Diversity and Evolution. In: eLS. John Wiley & Sons, Ltd; 2018: 1–20.
MeSH terms
Substances
Grants and funding
- 2023YFF1304900/Ministry of Science and Technology of the People's Republic of China
- 2024A1515013196/Science and Technology Department of Guangdong Province
- SLYJ2023B4004/Guangdong Forestry Administration
- GML2020GD0804/PI Project of Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou)
- GML2022GD0804/PI Project of Southern Marine Science and Engineering Guangdong Laboratory (Guangzhou)
LinkOut - more resources
Full Text Sources

