Genome-wide identification and gene expression pattern analysis of the glycoside hydrolase family 1 in Fagopyrum tataricum
- PMID: 39695944
- PMCID: PMC11654022
- DOI: 10.1186/s12870-024-05919-3
Genome-wide identification and gene expression pattern analysis of the glycoside hydrolase family 1 in Fagopyrum tataricum
Abstract
Background: The β-glucosidases (BGLU) of glycoside hydrolase family 1 hydrolyze the glycosidic bond to release β-D-glucose and related ligands, which are widely involved in important physiological processes in plants. Genome-wide analysis of the BGLU genes in the model crops Arabidopsis thaliana and Oryza sativa revealed that they are functionally diverse. In contrast, the BGLU gene family in Tartary buckwheat remains unclear.
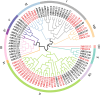
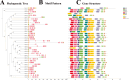
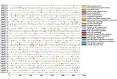
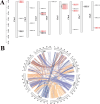
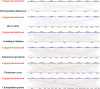
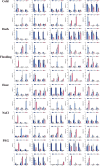
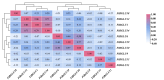
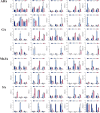
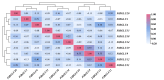
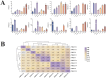
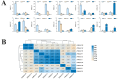
Results: This study identified the FtBGLU gene family based on Tartary buckwheat genomic data and analyzed the biological function of the FtBGLU gene using bioinformatics methods and the expression pattern of the gene using fluorescence quantitative PCR. The results showed that 39 BGLU genes were identified in Tartary buckwheat, which were classified into 10 subfamilies and one unclassified group. They were unevenly distributed on 10 chromosomes, and seven tandem duplication events involving 19 FtBGLU genes were observed, which mainly occurred in subfamily II. Their physicochemical properties are highly variable; however, they have relatively conserved exon-intron structures and high sequence homology in the subfamily, and most of the FtBGLUs contain conserved motifs, among which the expression products FtBGLU1, FtBGLU17, FtBGLU19, FtBGLU21, FtBGLU22, and FtBGLU28 have no β-glucosidase activity. Additionally, we analyzed the tissue expression specificity of 10 FtBGLU genes during Tartary buckwheat growth and development and their expression patterns under adversity stress and hormone treatments. Revealing the important role of the BGLU gene family in Tartary buckwheat growth and development, as well as its response to adversity, provides strong support for further analysis of its regulatory mechanisms and functional applications. A total of 39 FtBGLU genes were identified. Bioinformatics analysis of the gene structure, evolutionary relationship, and expression pattern of the Fagopyrum tataricum BGLU gene family establishes a foundation for a better understanding and future research on the Tartary buckwheat BGLU gene family.
Keywords: BGLU gene family; Fagopyrum tataricum; Evolutionary relationships; Expression patterns; Genome-wide analysis.
© 2024. The Author(s).
Conflict of interest statement
Declarations. Ethics approval and consent to participate: Plant materials, and collection do not necessitate licensing. The plant materials were maintained in accordance with the institutional guidelines established by the College of Agriculture at Guizhou University. The methodologies employed adhered to the pertinent guidelines and regulations. It should be noted that this study did not involve any human participants or animal experimentation carried out by the authors. Consent for publication: Not applicable. Clinical trial number: Not applicable. Competing interests: The authors declare no competing interests.
Figures

References
-
- Oliveira RP, Santos BV, Costal, Henrique MA, Daniel, Baffi MA. Xylanase and β-glucosidase production by aspergillus fumigatus using commercial and lignocellulosic substrates submitted to chemical pre-treatments. Ind Crop Prod. 2017;95:453–9. 10.1016/j.indcrop.2016.10.055. - DOI
-
- Gonzalez-pombo P, Farina L, Carrau F, Batista-Viera F, Brena B. A novel extracellular β-glucosidase from Issatchenkia terricola: isolation, immobilization and application for aroma enhancement of white Muscat wine. Process Biochem. 2011;46(1):385–9. 10.1016/j.procbio.2010.07.016. - DOI
-
- Opassiri R, Pomthong B, Akiyama T, Nakphaichit M, Onkoksoong T, Ketudat Cairns M, Ketudat Cairns JR. A stress-induced rice (Oryza sativa L.) beta-glucosidase represents a new subfamily of glycosyl hydrolase family 5 containing a fascin-like domain. Biochem J. 2007;408(2):241–9. 10.1042/BJ20070734. - DOI - PMC - PubMed
MeSH terms
Substances
Grants and funding
- Guida Lingjun Hezi (2023) 07/Research and Integrated Application of Key Technologies for Green and High Yield in Mountainous Characteristic Agriculture
- Qiankehe Platform Talent-BQW (2024) 009/Construction of Scientific and Technological Innovation Talent Team for High-Quality and High-Efficiency Mechanisation of Speciality Mixed Grains in Guizhou Province
- 32312051, 32160669, 32161143005/the Natural Science Foundation of China
LinkOut - more resources
Full Text Sources

