Screening and identification of potential target of 1'-acetoxychavicol acetate (ACA) in acquired lapatinib-resistant breast cancer
- PMID: 39698092
- PMCID: PMC11652900
- DOI: 10.1016/j.heliyon.2024.e40769
Screening and identification of potential target of 1'-acetoxychavicol acetate (ACA) in acquired lapatinib-resistant breast cancer
Abstract
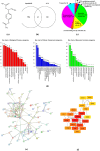
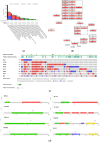
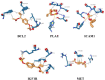
1'-Acetoxychavicol acetate (ACA) eliminates breast cancer cells via the HER2/MAPK/ERK1/2 and PI3K/AKT pathways, and it also directly influences endocrine resistance by both enhancing pro-apoptotic signals and suppressing pro-survival molecules. This study utilized bioinformatics to assess ACA target genes for lapatinib-resistant breast cancer. We identified differentially expressed genes (DEGs) using GSE16179 microarray data. DEGs from ACA-treated and lapatinib-resistant cells were analyses using Panther DB, Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analyses, and protein-protein interaction (PPI) network analysis. Genomic mutations, expression levels, prognostic significance, and ROC analysis were examined in selected genes. We used AutoDock Vina to conduct ACA molecular docking with potential target genes. In the PPI network analysis, BCL2, CXCR2, and CDC42 were the three highest-scoring genes. Genetic modification analysis identified PLAU and SSTR3 as the genes most frequently altered in breast cancer samples. The RTK-Ras pathway is likely to be affected by changes in BCL2, CXCR2, CDC42, SSTR3, PLAU, ICAM1, IGF1R, and MET genes. Patients with breast cancer who had lower levels of BCL2, SSTR3, PLAU, ICAM1, IGF1R, and MET had worse overall survival compared to other groups. ACA exhibited moderate binding affinity to BCL2, SSTR3, PLAU, ICAM1, IGF1R, and MET. Overall, ACA might counteract breast cancer resistance to lapatinib by targeting BCL2, SSTR3, PLAU, ICAM1, IGF1R, and MET. Further in vitro studies involving gene silencing could provide more detailed insights into the mechanism by which ACA combats lapatinib resistance.
Keywords: 1′-acetoxychavicol acetate; Bioinformatics; Breast cancer; Lapatinib-resistant; Targeted therapy.
© 2024 The Authors. Published by Elsevier Ltd.
Conflict of interest statement
The authors declare the following financial interests/personal relationships which may be considered as potential competing interests: Muhammad Da'i reports financial support was provided by 10.13039/501100016269Muhammadiyah University of Surakarta. If there are other authors, they declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Figures


References
-
- Jabbarzadeh Kaboli P., Salimian F., Aghapour S., Xiang S., Zhao Q., Li M., Wu X., Du F., Zhao Y., Shen J., Cho C.H., Xiao Z. Akt-targeted therapy as a promising strategy to overcome drug resistance in breast cancer – a comprehensive review from chemotherapy to immunotherapy. Pharmacol. Res. 2020;156 - PubMed
LinkOut - more resources
Full Text Sources
Research Materials
Miscellaneous

