Development of monoclonal antibodies against P. gingivalis Mfa1 and their protective capacity in an experimental periodontitis model
- PMID: 39699191
- PMCID: PMC11774036
- DOI: 10.1128/msphere.00721-24
Development of monoclonal antibodies against P. gingivalis Mfa1 and their protective capacity in an experimental periodontitis model
Abstract
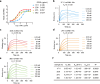
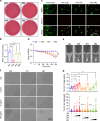
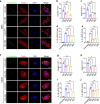
Porphyromonas gingivalis (P. gingivalis), a gram-negative, black-pigmented anaerobe, is a major etiological agent and a leading cause of periodontitis. Fimbriae protein Mfa1 is a key virulence factor of P. gingivalis and plays a crucial role in bacterial adhesion, colonization, biofilm formation, and persistent inflammation, making it a promising therapeutic target. However, the role of anti-Mfa1 antibodies and the underlying protective mechanisms remain largely unexplored. Here, we developed and characterized the monoclonal antibodies (mAbs) targeting the Mfa1 protein of P. gingivalis. Function analysis showed that anti-Mfa1 mAbs mediated bacterial agglutination and inhibited P. gingivalis adhesion to saliva-coated hydroxyapatite and host cells. Notably, anti-Mfa1 mAbs significantly reduced bacterial burden and alveolar bone loss in a P. gingivalis-induced experimental periodontitis model. These results show that anti-Mfa1 mAbs can be beneficial in alleviating P. gingivalis infections, and provide important insights for the development of adequate adjuvant treatment regimens for Mfa1-targeted therapeutics.
Importance: Fimbriae (pili) play an important role in bacterial adhesion, invasion of host cells and tissues, and formation of biofilms. Studies have shown that two types of fimbriae of Porphyromonas gingivalis, FimA and Mfa1, are important for colonization and infection through their binding to host tissues and other bacteria. While anti-FimA antibodies have been shown to improve periodontitis, the effect of anti-Mfa1 antibodies on P. gingivalis infection and periodontitis was previously unknown. In this study, we report for the first time that anti-Mfa1 monoclonal antibodies can reduce P. gingivalis infection and improve periodontitis. These findings suggest that Mfa1 represents a promising therapeutic target, and the development of anti-Mfa1 mAbs holds a potential as essential diagnostic and adjunctive therapeutic tools for managing P. gingivalis-related diseases.
Keywords: Mfa1; P. gingivalis; fimbriae; monoclonal antibodies; periodontitis.
Conflict of interest statement
The authors declare no conflict of interest.
Figures




Similar articles
-
Localization and function of the accessory protein Mfa3 in Porphyromonas gingivalis Mfa1 fimbriae.Mol Oral Microbiol. 2013 Dec;28(6):467-80. doi: 10.1111/omi.12040. Epub 2013 Oct 5. Mol Oral Microbiol. 2013. PMID: 24118823
-
Potent In Vitro and In Vivo Activity of Plantibody Specific for Porphyromonas gingivalis FimA.Clin Vaccine Immunol. 2016 Apr 4;23(4):346-52. doi: 10.1128/CVI.00620-15. Print 2016 Apr. Clin Vaccine Immunol. 2016. PMID: 26865596 Free PMC article.
-
Purification of Native Mfa1 Fimbriae from Porphyromonas gingivalis.Methods Mol Biol. 2021;2210:75-86. doi: 10.1007/978-1-0716-0939-2_8. Methods Mol Biol. 2021. PMID: 32815129
-
Molecular interaction of Porphyromonas gingivalis with host cells: implication for the microbial pathogenesis of periodontal disease.J Periodontol. 2003 Jan;74(1):90-6. doi: 10.1902/jop.2003.74.1.90. J Periodontol. 2003. PMID: 12593602 Review.
-
Porphyromonas gingivalis FimA and Mfa1 fimbriae: Current insights on localization, function, biogenesis, and genotype.Jpn Dent Sci Rev. 2021 Nov;57:190-200. doi: 10.1016/j.jdsr.2021.09.003. Epub 2021 Oct 7. Jpn Dent Sci Rev. 2021. PMID: 34691295 Free PMC article. Review.
Cited by
-
A Narrative Review of Periodontal Vaccines: Hope or Hype?Cureus. 2025 Mar 15;17(3):e80636. doi: 10.7759/cureus.80636. eCollection 2025 Mar. Cureus. 2025. PMID: 40091902 Free PMC article. Review.
References
-
- Dominy SS, Lynch C, Ermini F, Benedyk M, Marczyk A, Konradi A, Nguyen M, Haditsch U, Raha D, Griffin C, et al. . 2019. Porphyromonas gingivalis in Alzheimer’s disease brains: evidence for disease causation and treatment with small-molecule inhibitors. Sci Adv 5:eaau3333. doi:10.1126/sciadv.aau3333 - DOI - PMC - PubMed
MeSH terms
Substances
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
