ACSL4-mediated H3K9 and H3K27 hyperacetylation upregulates SNAIL to drive TNBC metastasis
- PMID: 39700137
- PMCID: PMC11670210
- DOI: 10.1073/pnas.2408049121
ACSL4-mediated H3K9 and H3K27 hyperacetylation upregulates SNAIL to drive TNBC metastasis
Abstract
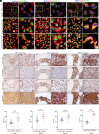
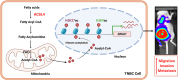
Triple-negative breast cancer (TNBC) has profound unmet medical need globally for its devastating clinical outcome associated with rapid metastasis and lack of targeted therapies. Recently, lipid metabolic reprogramming especially fatty acid oxidation (FAO) has emerged as a major driver of breast cancer metastasis. Analyzing the expression of major FAO regulatory genes in breast cancer, we found selective overexpression of acyl-CoA synthetase 4 (ACSL4) in TNBC, which is primarily attributed to the absence of progesterone receptor. Loss of ACSL4 function, by genetic ablation or pharmacological inhibition significantly reduces metastatic potential of TNBC. Global transcriptome analysis reveals that ACSL4 activity positively influences the gene expression related to TNBC migration and invasion. Mechanistically, ACSL4 modulates FAO and intracellular acetyl-CoA levels, leading to hyperacetylation of particularly H3K9ac and H3K27ac marks resulting in overexpression of SNAIL during the course of TNBC metastatic spread to lymph node and lung. Further, human TNBC metastasis exhibits positive correlation among ACSL4, H3K9ac, H3K27ac, and SNAIL expression. Altogether, our findings provide molecular insights regarding the intricate interplay between metabolic alterations and epigenetic modifications, intertwined to orchestrate TNBC metastasis, and posit a rational understanding for the development of ACSL4 inhibitors as a targeted therapy against TNBC.
Keywords: ACSL4; SNAIL; TNBC; histone acetylation; metastasis.
Conflict of interest statement
Competing interests statement:The authors declare no competing interest.
Figures






References
-
- Sung H., et al. , Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021). - PubMed
-
- Siegel R. L., Giaquinto A. N., Jemal A., Cancer statistics, 2024. CA Cancer J. Clin. 74, 12–49 (2024). - PubMed
-
- Gluz O., et al. , Triple-negative breast cancer–current status and future directions. Ann. Oncol. Off. J. Eur. Soc. Med. Oncol. 20, 1913–1927 (2009). - PubMed
MeSH terms
Substances
Grants and funding
LinkOut - more resources
Full Text Sources
Molecular Biology Databases
Research Materials

